One of the most important concepts in bioassay is that of replication. Test samples are regularly replicated 2 or 3 times. Multiple response measurements at each dose will, in general, result in a better estimate of the precision of our result.
As with all biological experiments, however, there is often quite a lot of variability in replicates (unless they are pseudo replicates. Read our blog on the subject to find out more: Understanding true replicates and pseudo replicates in bioassay) and we often see replicates which show noticeably different responses for the same dose.
Unless there are known causes for this from some laboratory mishap, many clients feel the need to make decisions about what level of variability within replicates is acceptable. At Quantics, one metric we often see clients using to make such decisions is the coefficient of variation, or CV. This is defined as the ratio of the standard deviation and the mean of the replicates, and is often expressed as a percentage (often refered to as %CV).
If the CV is being used in this way, then the suitability criterion on replicate variability is typically considered valid if the CV of all replicate sets falls below a chosen value. So, for example, an assay might be considered good to go if all of five sets of replicates at five different dose levels have a CV of below 20%.
Key Takeaways
-
The coefficient of variation (CV) is calculated as the standard deviation divided by the mean of replicates. Although many use it to judge assay repeatability, its value inherently changes with the dose because the mean response varies, even when the standard deviation is constant.
-
Since homogeneity of variance implies a constant standard deviation across doses, a the CV will vary naturally with the mean. This means a fixed CV cut-off applied uniformly across doses can be misleading, as lower mean responses (or higher ones, depending on assay direction) will automatically produce higher CVs, regardless of true data variability.
-
Some mistakenly use CV as an outlier detection tool. Since CV is independent of the model fit, however, it does not capture whether data points are anomalous relative to a fitted model. This disconnect makes it a poor criterion for assessing overall assay suitability.
The standard deviation of the replicates is just a property of the replicates and is independent of the model fit. It is calculated by, for each dose,
- Measuring the distance of each replicate point from the mean of the points,
- Square each distance, and add up
- Divide by n-1
- Take the square root.
(If you want to know why then give us a call!)

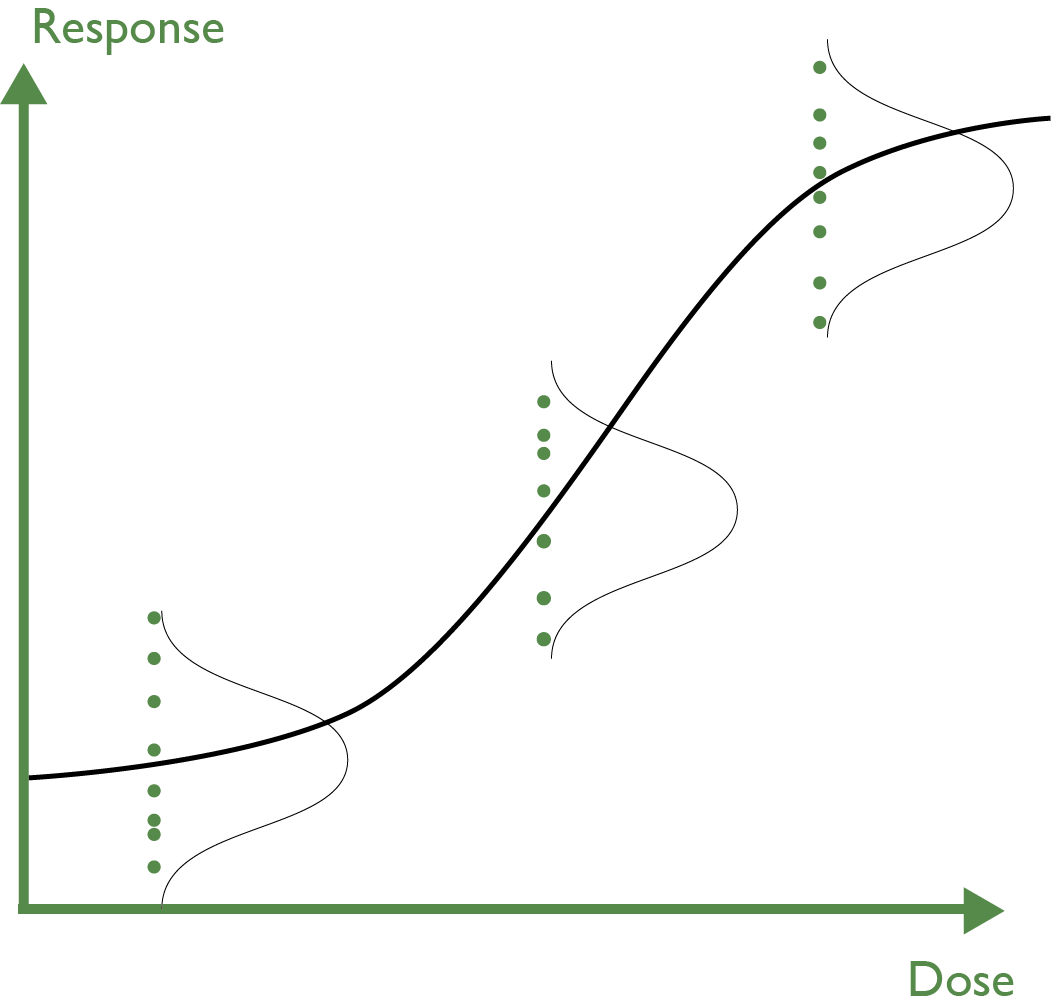
Now let’s consider the analysis. If the analysis is going to provide confidence intervals, or you plan to use any suitability criteria that take the variability of the data into account (such as F-tests or equivalence tests), then there is a built-in assumption that the standard deviation of the replicates at each dose is the same. This is called homogeneity of variance, and applies to any model we choose to fit.
If the standard deviation is not constant across the doses, then any inferences we might make for suitability criteria will not be valid. (A log transform of the response, for instance, may achieve homogeneity of variance in a case where we have standard deviation increasing with dose. Read When and How Should I Transform my Response Data? to find out more.)
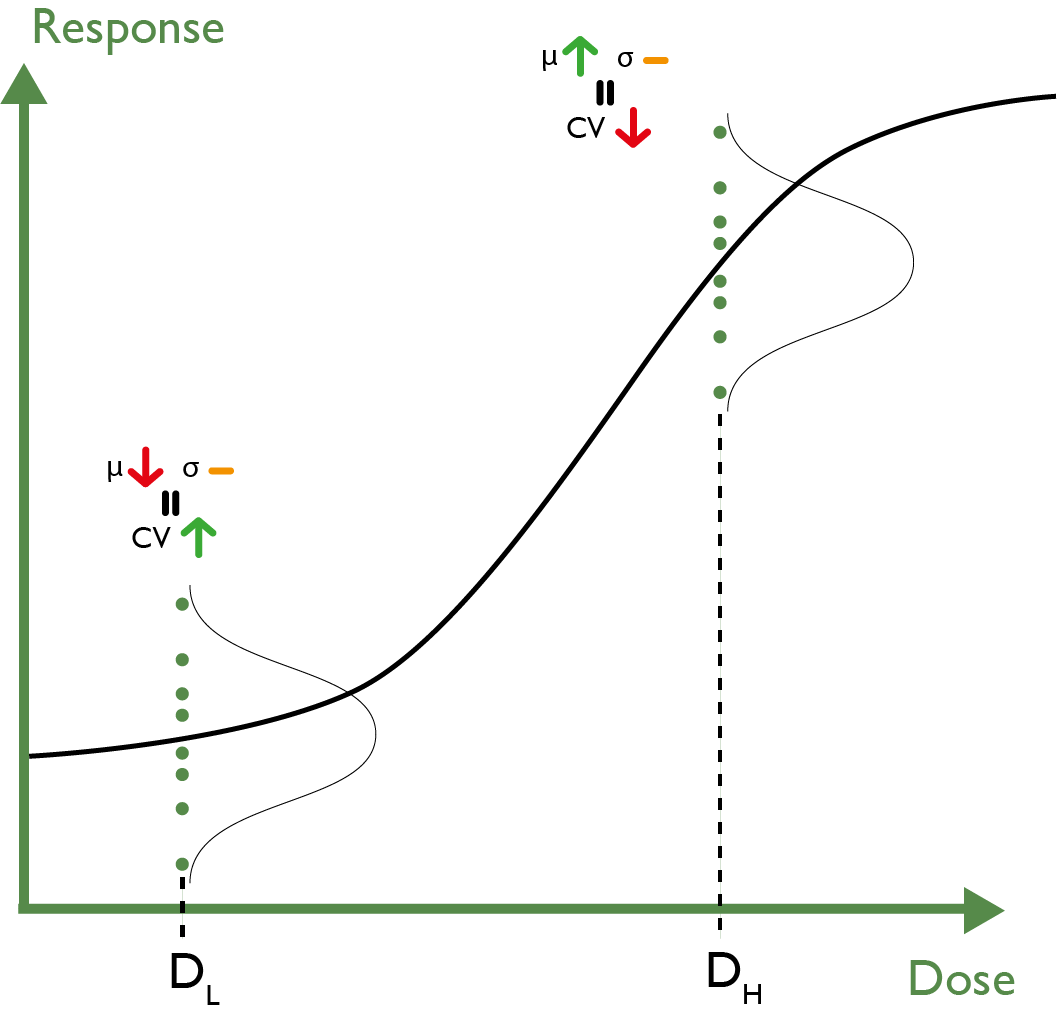
This requirement for a constant standard deviation leads to the downfall of a constant CV cut-off. Recall the definition of CV: standard deviation divided by mean. If we have a dose response, then the mean of the replicate distributions will change with dose, but—if we have the required homogeneity of variance—the standard deviation will be constant across the doses. This means that for a positive slope assay, the replicate CV should decrease across the dose range, and vice versa for a negative slope assay.
Using a CV cut-off that is the same for each dose, therefore, makes no sense.
The only time we would expect to see a constant CV of the raw responses is when the responses need a log transformation to achieve homogeneity of variance. But in that case, basing suitability criteria on the raw scale makes no sense either, as these are not the numbers we will be using to calculate the reportable value.
Alongside those who use CV as a repeatability criterion, we also find that some clients think of CV as a kind of outlier measure. This is again flawed reasoning. Remember that the CV of a set of replicates is independent of the model fit, so it tells you nothing about whether the data might be considered an outlier with respect to the model.
Quantics, therefore, strongly advises against using CV as a suitability criterion, whether for assay repeatability or outlier management. Our team of statisticians are on hand to help you manage these and any other statistical aspect of your bioassay—just give us a call!


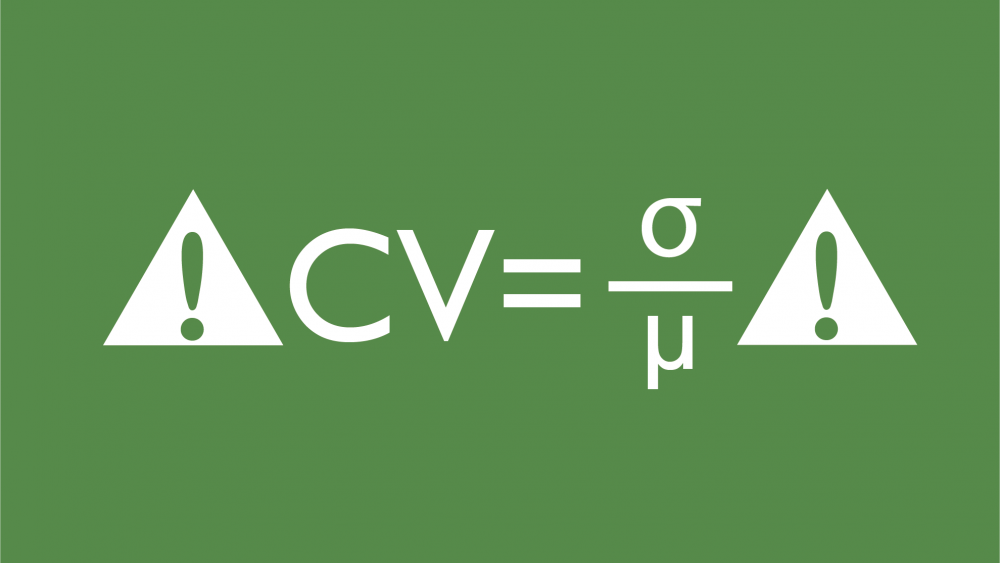



Comments are closed.