Pseudo-replicates and Bioassay
Here, we investigate the difference between true replicates and pseudo-replicates, and discuss the problems that may arise if this difference is ignored or misunderstood in bioassay statistical analysis.
Quality Control by sampling
Consider a manufacturing process that makes widgets. To check the quality of a batch, a number of widgets randomly selected from the batch can be examined in detail to draw conclusions about the batch as a whole. You might perhaps select 100 from each batch of 1000. This is standard QC practice. In general, the more widgets you sample, the greater the certainty that the batch is acceptable.
But this only works if the samples are independent. Because they are sampled randomly, there is no reason to think that one widget’s test result is related to another’s.
Key Takeaways
- True replicates come from independent samples, while pseudo-replicates (technical replicates) measure the same sample multiple times. Confusing the two leads to incorrect statistical inference in bioassay analysis.
- Naively using pseudo-replicates for calculations underestimates variability, producing overly narrow confidence intervals and misleading suitability results. This may cause unnecessary assay failures—or falsely pass poor-quality samples.
- Pseudo-replicates help measure plate variability during assay development, but are generally inefficient in commercial bioassays. Removing unnecessary pseudo-replicates can significantly improve assay throughput.
But if you selected only 10 widgets and tested each one 10 times, you would still obtain 100 measurements—but this is clearly less reliable. This is pseudo-sampling or pseudo-replication. The samples tested are not independent, and results are expected to be almost identical because they come from the same widget. These are sometimes called technical replicates.
Pseudo-replicates in bioassay
This issue is very common in bioassays. Assays are designed to test samples from a batch for QC purposes. Testing more independent samples increases confidence in the result. But how the experiment is run makes all the difference.
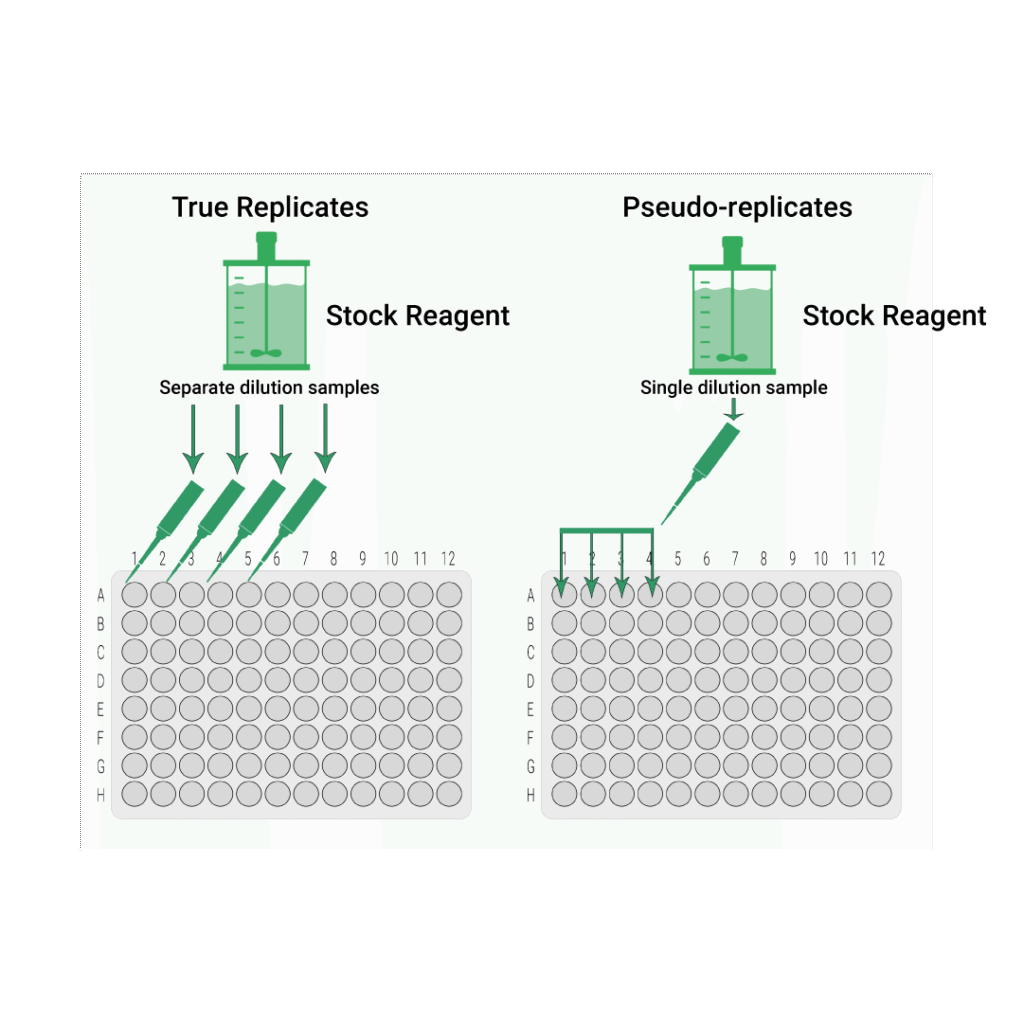
If a single sample is taken from the batch, diluted, and then pipetted into adjacent wells, this creates multiple measurements of the same sample. The results are expected to be nearly identical. Differences reflect only plate variability, not sample-to-sample biological variability.
Why is this a problem?
The problem arises when statistics are applied. In general, bioassay statistical tests assume independent samples. Using pseudo-replicates as if they were independent produces incorrect, usually artificially narrow, confidence intervals.
This underestimation of biological variability can lead to incorrect suitability outcomes—either rejecting good assays or passing assays that should fail. Even if scientists do not explicitly calculate confidence intervals, any suitability criterion that depends on variability will be affected.
The F test for goodness of fit
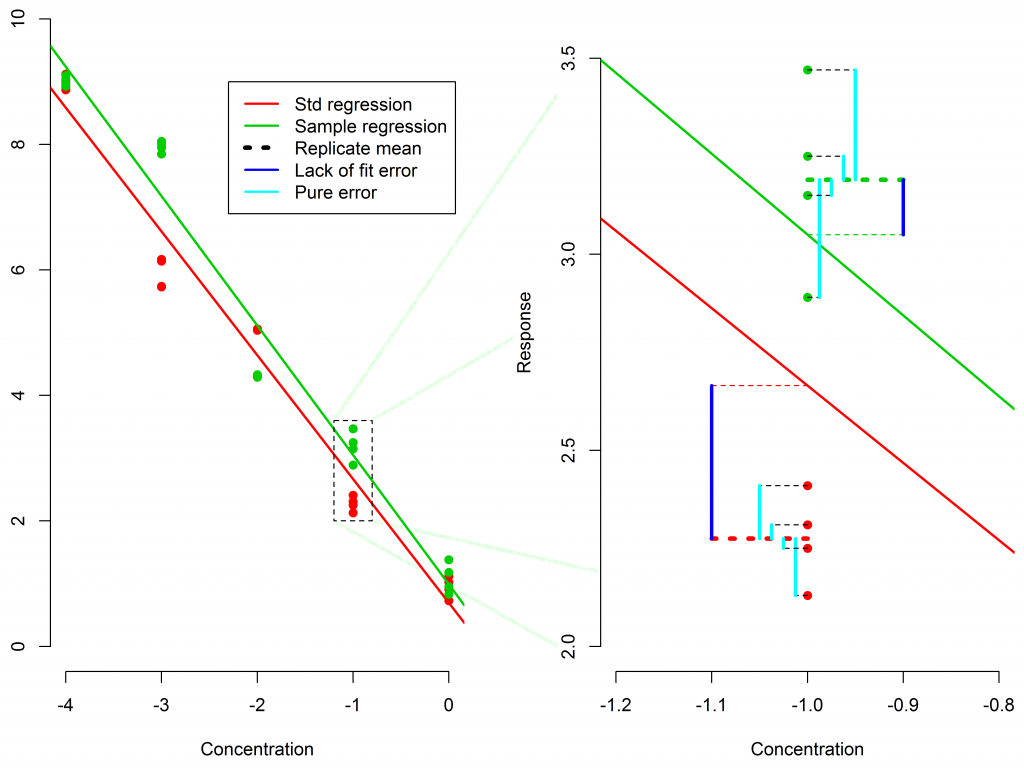
The F test evaluates whether the variability of the mean replicate responses around the model fit (lack-of-fit error) is significantly larger than the variability of the replicates around their mean (pure error).
Pure Error measures how much the replicates differ from their own mean.
Lack of Fit Error measures how far the replicate means are from the fitted model.
If lack-of-fit exceeds pure error, the goodness-of-fit may fail depending on the threshold.
But with pseudo-replicates, pure error is artificially tiny, because the assay is measuring the same sample repeatedly. This makes the F-test extremely sensitive, often causing failures even when the curve is perfectly acceptable.
So why use pseudo-replicates at all?
Pseudo-replicates are useful during development to measure within-plate variability. In these cases, the replicates are treated as if they were separate samples purely to quantify technical noise.
Outside of development, however, pseudo-replicates usually waste valuable plate space without contributing meaningful information about sample-to-sample variation.
As assays move from development to commercial use, where throughput and efficiency matter, it becomes increasingly important to decide whether pseudo-replicates are needed at all.








