Should I Average My Pseudo-replicates?
A common discussion we at Quantics have with our clients surrounds the issue of pseudo-replicates. It’s an area of tension between scientist and statistician – pseudo-replicates reduce the cost of an assay, but also its statistical validity, meaning that a compromise solution is often required. One such solution is averaging pseudo-replicates, which comes with its own set of pros and cons. We want to examine when pseudo-replication might prove problematic and take an empirical look at why averaging your pseudo-replicates may be a good idea.
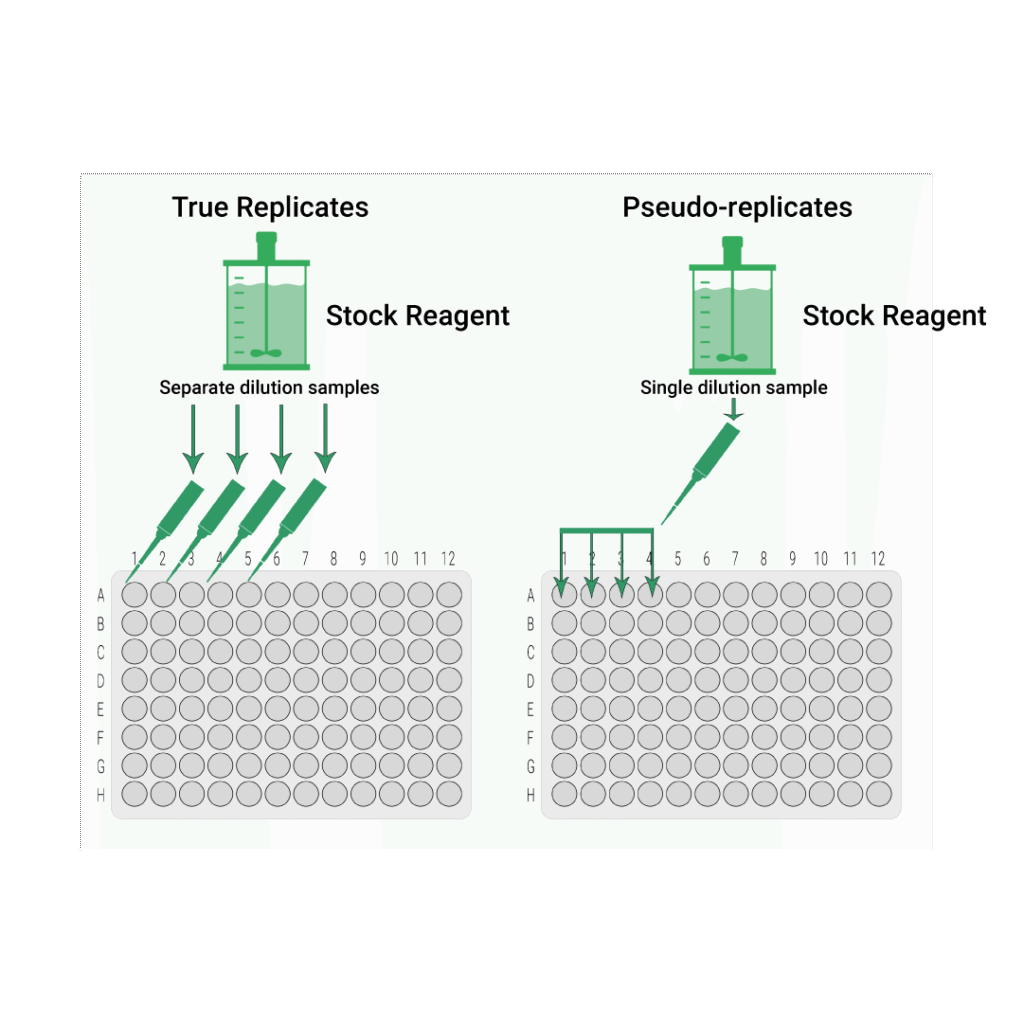
True Replicates vs Pseudo-replicates
In bioassays, there are two main ways of making up a set of dilution series from a stock solution of our lot.
True Replicates: Each replicate dilution series is made up from a separate draw from the stock solution.
Key Takeaways
- Pseudo-replicates introduce correlation between responses at a dose, breaking the independence assumption that underpins confidence intervals and F-tests, and even modest correlations can seriously distort those results.
- Averaging pseudo-replicates at each dose level removes that correlation and brings the behaviour of confidence intervals and parallelism tests back in line with their expected error rates in simulation studies.
- Averaging all replicates sacrifices information on between-replicate variability and prevents some tests (such as goodness-of-fit F-tests), but it simplifies analysis and guards against hidden correlations in routine bioassay work.
Pseudo-replicates: All replicate dilution series for the sample are made up from a single draw from the stock solution of that sample.
To understand this further, let’s imagine you are charged with serving a number of glasses of orange juice to be prepared from a packet of juice concentrate. As before, there are two ways you can do this. You could put a small amount of concentrate in each glass and then dilute by adding water. This would be similar to the idea of true replicates in an assay. Alternatively, you could put the concentrate in a jug, dilute, and fill each glass from the jug. This would be like using pseudo-replicates.
The glasses of juice prepared using the first method will have far more variable strengths than those prepared using the second method. No matter how careful you are, there will always be subtle differences in the amount of concentrate and the amount of water added to each glass. By contrast, since the dilution step only occurs once in the second method, this variability does not occur. And, indeed, pseudo-replicates are often significantly less variable than true replicates when used in bioassays since variability associated with sampling and dilution are not included.
We can also see how the juice strength across the glasses is correlated when using the second method – indeed, if the jug of juice was perfectly mixed, we would expect the strengths to be identical. We can easily predict the strength of all the glasses of juice just by tasting one.
Using the first method, however, the juice strength in one glass is independent of the strength in another glass. Adding a little bit more water to one glass has no effect on any others and we gain little information about the strength of the juice in other glasses when we taste just one. We see similar effects with replicates in bioassays: true replicates are independent, while pseudo-replicates are correlated.
The Problem with Psuedo-replicates
At Quantics, we often see our clients using pseudo-replicates in their bioassays. From the perspective of a scientist in a working laboratory, this makes sense: each sample preparation requires only one draw from the stock solution. This means that stock is used more efficiently, saving resources in the short term. And an assay with reduced variability is good, right?
Not so fast. From the perspective of a statistician, pseudo-replication introduces some serious problems in the analysis of the assay and the conclusion it draws. The key problem is the presence of correlation among the responses. When we perform statistical inference – that is, calculate a confidence interval or use a statistical test that returns a p-value – a fundamental assumption is that the responses are independent. Clearly, this assumption does not hold when using pseudo-replicates since they often exhibit significant correlation.
A Simulation Experiment
How much of a problem is this? We examined the effect of pseudo-replication using simulated bioassays. For each assay, reference and test samples were simulated from parallel four parameter logistic (4PL) curves with a known relative potency. At each dose level, two replicates were simulated such that correlation of the duplicates ranged from 0 (independent replicates) to a very strong correlation of 0.95.
To determine the effect of the correlation on statistical inference, there were three criteria examined:
95% confidence interval on the relative potency: How often does the true relative potency fall within the 95% confidence interval (CI) for the relative potency of the simulated assays? If our inferential procedure is performing as expected, this should be the case for about 95% of the simulated assays. We would consider an assay to have “failed” on this metric if the 95% CI does not contain the true RP, and to have “passed” when the 95% CI contains the true RP. So, we would expect to see a 5% “failure” rate for a 95% confidence interval.
Goodness-of-fit F-Test at the 5% level: The data were simulated using a 4PL model, so a 4PL model should fit the data well. However, we expect that a percentage of assays will fail by chance due to random variability. If our hypothesis test is behaving as expected, then (approximately) 5% of simulated assays should fail – that is, return a p-value of less than 0.05 – by chance.
Parallelism F-Test at the 5% level: The assays were simulated such that the reference and test models were parallel (i.e., their A, B and D parameters were simulated to be identical). As with the goodness-of-fit test, we should see (about) 5% of parallelism tests on the simulated assays fail by random chance when the test is behaving as expected.
In total, 6250 assays were simulated. The results of these simulations against the test criteria are listed below:
| Correlation of Replicates | Failure rate for 95% CIs | Failure rate for GoF Test | Failure rate for Parallelism |
| Independent: True Replicates | |||
| 0 | 4.20% | 4.0% | 4.1% |
| Correlated: Pseudo-Replicates | |||
| 0.25 | 9.80% | 17.3% | 10.4% |
| 0.5 | 11.00% | 43.0% | 15.4% |
| 0.75 | 15.50% | 84.1% | 27.6% |
| 0.95 | 22.70% | 99.6% | 35.7% |
With independent replicates, the simulations show that all the tests were behaving as we might expect: close to 5% of assays failed on 95% CIs, while both F-Tests failed at close to the expected 5% rate. The presence of correlation, however, had a significant impact on the performance of these statistical inference methods. The goodness-of-fit F-Test was the most sensitive, with assay failure almost guaranteed at the strongest correlation. Even with low correlation of 0.25, almost 20% of assays failed the goodness-of-fit F-Test, much higher than the expected 5%.
The parallelism F-test was less sensitive, but still showed elevated failure rates with correlated replicates. With low correlation of 0.25, about 10% of assays failed the test, while 35.7% of assays failed when the correlation reached 0.95. Similarly, the 95% CI failure rate rose to near 10% when correlation was 0.25, and was as high as 22.7% with correlation of 0.95.
Pseudo-replicates in real assays may not exhibit correlations as strong as those at the high end of those used in the simulations – indeed, it is difficult to determine exactly how correlated replicates are. Nevertheless, these results demonstrate just how risky it can be to treat pseudo-replicates as independent. Even weak correlations can cause problems when not accounted for.
An Average Solution
The good news is that there are methods to account for the presence of correlation among pseudo-replicates (or any other replicates for which a significant degree of correlation is suspected). The simplest is to average the pseudo-replicates. That is, for each dose level, the mean (either arithmetic mean or geomean, depending on whether the response is on a linear or log scale, respectively) of all pseudo-replicate responses is found. This mean is then treated like a data point for model fitting.
Averaging removes the impact of the presence on correlation on the analysis: you cannot have correlations between replicates at a dose level if there’s only one response!
This comes at a cost, however. Averaging means we have only a single data point at each dose level, meaning we lose information about between-replicate variability. Some statistical tests also require multiple replicates to work – most notably the F-Test for goodness-of-fit.
To determine empirically the effect of averaging, a second simulation was performed using the same data analysed before but where we averaged the dose replicates before fitting the 4PL models and performing any statistical inference. As mentioned previously, it is not possible to perform a goodness-of-fit F-Test using an average of the replicates, so this test was omitted from this phase of the experiment. The results are shown below:
| Correlation of Replicates | Failure rate for 95% CIs | Failure rate for GoF Test | Failure rate for Parallelism | |||
| Replicates | Averaged | Replicates | Averaged | Replicates | Averaged | |
| Independent: True Replicates | ||||||
| 0 | 4.20% | 3.40% | 4.0% | — | 4.1% | 4.4% |
| Correlated: Pseudo-Replicates | ||||||
| 0.25 | 9.80% | 6.80% | 17.3% | — | 10.4% | 4.7% |
| 0.5 | 11.00% | 4.20% | 43.0% | — | 15.4% | 3.9% |
| 0.75 | 15.50% | 4.60% | 84.1% | — | 27.6% | 5.4% |
| 0.95 | 22.70% | 6.20% | 99.6% | — | 35.7% | 2.6% |
There was a marked improvement in the performance of the statistical inference methods when the averaged replicates were used. The confidence interval for the RP behaved as we would expect with even the strongest correlations, as the assay failure rate on 95% CIs remained close to 5% for all strengths of correlation. Additionally, a far smaller percentage of assays failed a parallelism F-Test when replicates were averaged.
So, should I average my pseudo-replicates – or even all my replicates?
The results of these statistical experiments make clear that correlations between replicates can significantly impact any statistical inference that is performed on the data. They also demonstrate that averaging replicates is an effective way addressing the impact of the correlation and ensuring that statistical inference procedures behave as expected. It would, therefore, be prudent to average pseudo-replicates before conducting any statistical modelling. The preparation of pseudo-replicates all but guarantees there will be some degree of correlation among the replicates and, while these correlations may not necessarily be strong, the results show that even weak correlations can cause problems for some common tests.
It is an interesting question whether this averaging should be extended to all replicates, regardless of whether they are believed to be true replicates or pseudo-replicates. Independent replicates performed well when used in statistical inference whether averaged or not, so there is an argument that a simplified procedure of averaging all replicates could be useful. The difference between true replicates and pseudo-replicates is nuanced, and to have different procedures for the two cases could conceivably lead to problems when the two are confused. It is also easily possible to have correlations between replicates which have been prepared as true replicates due to, for example, plate effects. These are unlikely to be predictable in advance like correlations due to pseudo-replication, so a policy of averaging all replicates can prevent the unknown introduction of errors.
There are, however, significant downsides to averaging replicates. We’ve mentioned these previously: averaging means we lose all information about between-replicate variability, which is useful when determining replication strategy. Some statistical tests also require multiple replicates at each dose level to be used, notably the goodness-of-fit F-Test.
There are also other statistical methods available to make use of correlated data without the need for averaging, one such being mixed effect models. These account for the correlation structure of the responses in the model, meaning independence is no longer required for statistical inference. This has the advantage that correlated data can be used without losing information about between-replicate variability. Mixed models, however, are more complicated and require sophisticated bespoke statistical software to be implemented. This contrasts with averaging replicates, which is a simple process that can be handled by most off-the-shelf software.
Pseudo-replication is not an inherently bad idea: there are many circumstances in which it might be a good solution for an assay. Nevertheless, the results of the experiment outlined here demonstrate clearly that pseudo-replicates must be treated with care thanks to correlations. Averaging your pseudo-replicates is, therefore, an important step to ensure that statistical inference processes in an assay behave as expected.
This blog is based on work completed by Liam Cann in support of his MSc in Statistics at the University of New Brunswick under the joint supervision of Dr Connie Stewart and Quantics Director of Statistics Matthew Stephenson.