What is Parallelism in Bioassay?
In the most basic sense, relative potency bioassays consist of a comparison of the behaviour of a reference standard to that of a test sample. That process is not the topic of discussion here (we’ve covered it extensively elsewhere, in Potency Assay: Relative or Absolute, for example). What we want to examine is one of the foundational assumptions of the relative potency calculation: parallelism.
Models, Models, everywhere
Parallelism is important for all models used in relative potency bioassay, but to keep things simple we’re going to focus on just the 4PL curve for now.
We described this model in detail elsewhere (See Complications of fitting 4PL and 5PL models to bioassay data, What is the 4PL Formula?), but, broadly, it is a “S-shaped” (or sigmoidal) curve. It consists of flat asymptotes at high and low doses with a steeper transition region at intermediate doses.
Key Takeaways
- In relative potency bioassays, parallelism (or similarity) is the assumption that the dose–response curves of a reference standard and a test sample are essentially horizontal translations of each other. This condition is key to defining an unambiguous measure of relative potency.
- If the curves are non-parallel, then the test sample does not behave as a dilution of the reference. This means the horizontal shift varies with the dose, making the relative potency measurement ambiguous.
- There are two main methodologies for assessing parallelism. Significance testing compares an unconstrained model with a constrained model using a statistical test, such as the F or \( \chi^2 \) tests. Equivalence testing uses pre-defined equivalence limits based on historical data to determine if the observed non-parallelism is within acceptable bounds.
We say the high-dose (properly infinite-dose) asymptote is defined by the A parameter and the low-dose (zero-dose) asymptote is defined by the D parameter. Then, the B parameter encapsulates the slope of the transition region (though is not equal to it), and the C parameter relates to the EC50, which is the dose at which half the maximal response is observed. These can be seen in Figure 1.

Ok, but how can curves be parallel?
For the time being, we’re going to assume a perfect maths world where statistical variability does not exist. This allows us to talk about, for example, model parameters being equal. We’ll reintroduce the real world when we tackle statistical parallelism testing later!
Parallelism is a classic example of when the everyday meaning of a word differs from its use in a technical context. When we talk about lines being parallel in, say, high school geometry, we mean straight lines which will never cross. Formally, geometrically parallel straight lines have the same gradient.
Parallelism is a classic example of when the everyday meaning of a word differs from its use in a technical context. When we talk about lines being parallel in, say, high school geometry, we mean straight lines which will never cross. Formally, geometrically parallel straight lines have the same gradient.
In that context, however, it is not immediately clear what it would mean for curves—such as the 4PL and 5PL—to be parallel. To answer that question would certainly go beyond the high school geometry textbook!
The solution—in bioassay at least—is similar to the requirement that the gradients of parallel lines be equal: namely that a subset of the available parameters be equal in two parallel models. For the 4PL, the A, D and B parameters must all be equal, with only the C parameter allowed to differ. That is, the models must be horizontal translations of each other. This can be seen in Figure 2.
Why do we care about parallelism?
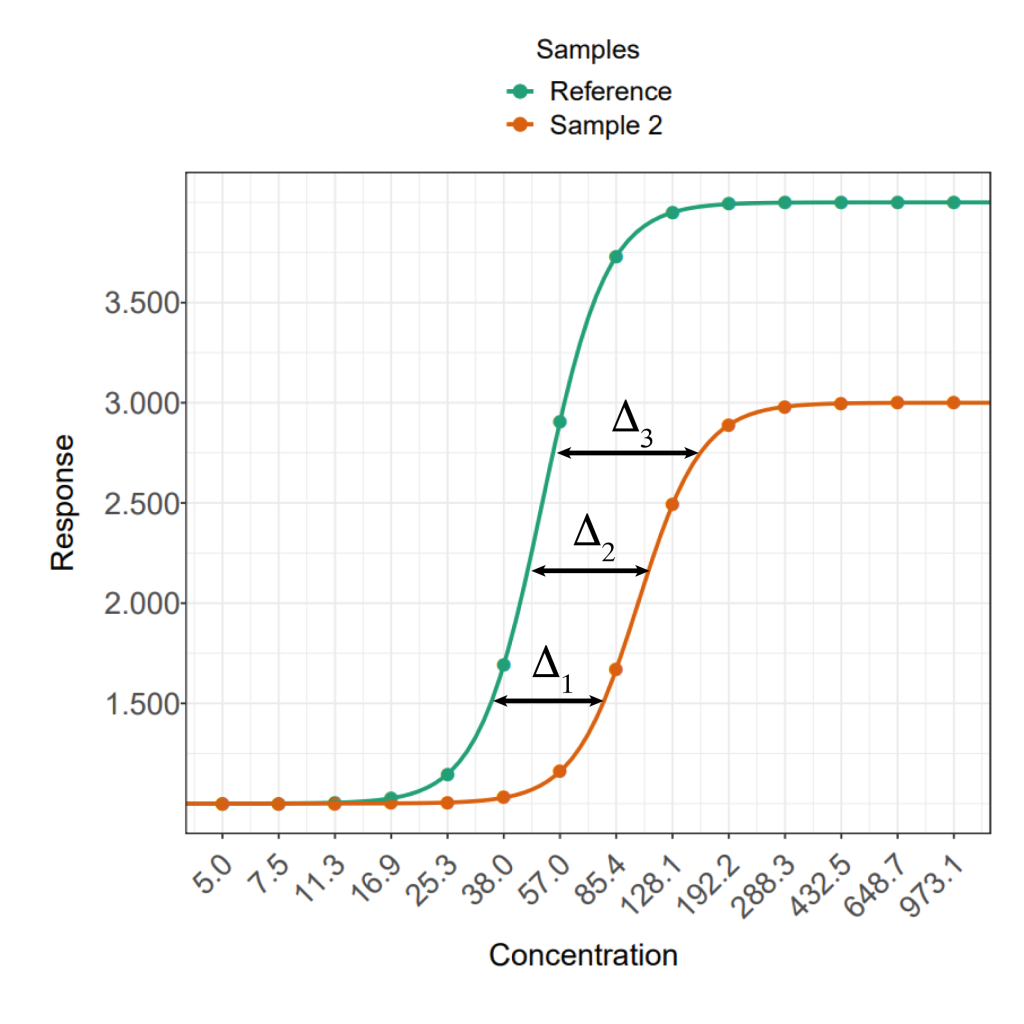
In many biological reactions, as the dose of reagent increases, the response remains near-constant at a low level, before increasing quickly and saturating at a near-constant high level. That’s why 4PL models are used to model bioassay data: the dose-response relationship of many reactions seen in bioassays follow that same S-shaped pattern.
Importantly, for two samples of the same reagent, the response at the near-constant plateaus and the rate of change of the response between them will be the same. If one of the samples is more diluted than the other, then the response for a given concentration will be lower, resulting in a shift to the right. Those important touchpoints—near-constant plateaus and rate of change—will, nevertheless, remain identical.
Sound familiar? We can model the dose-response curves of dilutions of the same reagent with parallel curves! The samples would be well fit by models with equal A, B, and D parameters, with only the C parameter potentially differing. In that sense, we say those dose-response relationships are themselves parallel, and the assay has passed parallelism.
Conversely, if any of the A, B, or D parameters of the two models were not equal, this is a sign that the test sample is not behaving as a dilution of the reference. We would say that these data are non-parallel, and the assay had failed parallelism.
For a relative potency to be well defined, we require the data for the two samples to be parallel. That’s because the relative potency of a test sample is how far its dose-response curve is shifted along the x-axis compared to that of the reference. For parallel data, this distance is constant with dose in the non-asymptotic regions of the curve. For non-parallel data, however, it can vary, meaning the relative potency is ambiguous.
This is why parallelism matters: it means you can measure one, unambiguous value for the relative potency of a test sample. You could define a similar quantity to relative potency for non-parallel data, but it would vary with the response level and be nowhere near as useful.
Parallelism Testing
This is where the real world re-enters the picture. Biological experiments are complex and highly variable. Even when we compare dilutions of the same reagent, we are unlikely to get datasets which are truly parallel in the formal, mathematical sense. To counter this problem, we implement statistical tests for parallelism to determine whether the degree of non-parallelism we observe is due to statistical chance or the two samples truly behaving in different ways.
Our paper Parallelism in Practice: Approaches to Parallelism in Bioassays outlines parallelism testing in great detail using simulated data. For now, we’re going to give a quick overview of the available options.
There are, broadly, two philosophies when it comes to parallelism testing (and other suitability criteria, for that matter). One question we can ask is “are the two datasets significantly non-parallel?”. This methodology is known as significance testing. It assumes the datasets are parallel (formally, the null hypothesis (H0) is the data are parallel), and checks the strength of the evidence against this (the alternative hypothesis (H1) is the data are non-parallel).
A different question we could ask is “is the non-parallelism we observe enough to be problematic?” This is equivalence testing, and assumes non-parallelism (H0: data are non-parallel) while checking if the data are actually parallel enough to be useful (H1: data are parallel).
Let’s take a deeper look at these two methodologies.
Significance testing
When we perform a significance test, we first fit a model as normal to our test and reference datasets separately. This is known as the unconstrained fit: the models for test and reference data need not have any of the same parameter values.
The next step is to fit a constrained model. This time, the model fitted to the test data is forced to be parallel to the reference model—recall: this means the two curves have the same A, B and D parameters.
The quality of these two fits is then compared. The constrained fit will always be worse than the unconstrained since at least 3 of the model parameters are deliberately non-optimal. The question is how much worse.
This comparison uses a quantity known as a p-value. In the case of parallelism, the p-value tells us the probability of two truly parallel datasets seeming to be non-parallel only through random variability in the data. The p-value is generated by modelling the statistical variation of the data in the experiment, usually using either the F or \( \chi^2 \) distributions. Depending on which is used, the resulting test would be called a F test or \( \chi^2 \) test respectively (See Failing Goodness of Fit – How to Combat the F-test Headache and R² II: What should you use to measure Goodness-of-Fit? for more about the F test).
To use the p-value, we set a significance level (usually 0.01 or 0.05). This is the point below which the probability of these datasets seeming non-parallel through random variation alone is so small that we have to accept that they’re just non-parallel. If the p-value is smaller than the significance level, therefore, the assay has failed the parallelism test. Otherwise, we have no evidence that the datasets are truly non-parallel, so we treat them as parallel.
Significance testing is a very simple method for parallelism testing. The distributions used are readily available in off-the-shelf statistical software packages, meaning it is essentially plug-and-play. There’s very little assay-specific work needed to use significance testing.
There are, nevertheless, some well-known pitfalls. Significance tests are a binary measurement of parallelism: they tell you whether two datasets are parallel or not. They don’t give any information about how non-parallel those datasets are. They also compare all three constrained parameters at once, which means you don’t know which of them is causing the non-parallelism.
Significance tests also perform poorly in extremely precise or extremely variable assays. We’ve covered the former case in Failing Goodness of Fit – How to Combat the F-test Headache . In the latter, the variability of the assay can, effectively, “mask” the non-parallelism. Recall that the p-value was the probability of the datasets being non-parallel by random variability alone. If there is a lot of random variability, then the datasets would have to be very non-parallel before they generate a low enough p-value to be detected. This could allow datasets which probably wouldn’t give meaningful relative potency values to slip through the cracks.

Equivalence testing
Equivalence testing operates on the principle that we should only be worried about non-parallelism when it becomes problematic for the operation we are concerned with (i.e. estimating the relative potency). It takes this into account by using equivalence limits. These are values outside of which we consider our test measure as an issue.
To perform an equivalence test, we usually compare one of the constrained parameters of the reference model to that of the test model. For example, we could choose to look at the difference between the A parameters of the models.
We then set our equivalence limits for a 90% confidence interval (90% CI) for that difference. This is a statistically involved procedure which requires a lot of information about the behaviour of the assay, usually in the form of historical data. We cover the process of setting equivalence limits in How to Set Suitability Criteria for Relative Potency Bioassays.
For the equivalence test, we calculate the 90% confidence interval (90% CI) on our chosen measure of parallelism. Recall that the null hypothesis of an equivalence test is the chosen measure is problematic: it falls outside of the equivalence limits. If any of our 90% CI falls outside of the equivalence limits, we do not have enough evidence to reject the null hypothesis as the probability of the value being within the acceptable limits is too low. This means the assay fails parallelism.
Conversely, if the entirety of the 90% CI is within the limits, we have sufficient evidence to conclude that the non-parallelism we observe is not a problem, and the assay passes parallelism.
Note that the 90% CI doesn’t have to be centred on or even include the value of our measure of parallelism which would indicate true parallelism (e.g. 0 if using the difference of A parameters) in order to pass an equivalence test. Since the inclusion of this value is equivalent to passing a significance test, this means that assays which fail such tests can pass an equivalence test.
Equivalence testing also avoids some of the pitfalls of significance testing we mentioned earlier. It does not penalise overly precise assays, since the only effect of high precision is to narrow the 90% CI. If anything, this makes very precise assays more likely to pass! Similarly, the masking of non-parallelism in highly variable assays is likely to be avoided since the 90% CI will be widened, meaning it is more likely to fall outside of the equivalence limits.
The major downside of equivalence testing is the amount of historical data required to set meaningful equivalence limits. This means that its unlikely to be of use for early assay development, since that data simply may not exist.
Parallelism in Practice
As we’ve outlined, parallelism is probably the most important suitability criterion to keep track of for relative potency bioassays. If nothing else, relative potency is meaningless without the assumption of parallelism! That’s why we recommend stringent parallelism testing to be a part of every assay lifecycle.
Which testing methodology to use? Both have their benefits and drawbacks as we’ve mentioned. Both are mentioned in regulatory guidance too: the USP recommend the use of equivalence testing, while PhEur recommend significance testing and accept equivalence testing. General opinion among the statistical community seems to be moving in the direction of equivalence testing being the preferred option.
As always, however, every assay will have its own nuances, which means the right combination of choices will be different for each one!
If you have any questions about parallelism for your bioassay, don’t hesitate to reach out: https://www.quantics.co.uk/contact-us2/