Bioassay Simulation: Reduce Lab Work
Bioassay simulation can be used during assay development maintenance to supplement and complement the information gained from lab experiments.
Bioassay simulation assumes that the available real data is a representative sample of all the real data sets that might be created over the life of the assay. The mathematical properties of this sample can be used to create a range of new data sets with previously untested but realistic characteristics, spanning the complete range of data sets and doses that might be expected to be seen or tested in real use.
These new data sets can be created at any dose (or dilution) and can be used to guide the choice of doses to use, the number of replicate wells needed and even the impact of different plate layouts so that, if further lab work is needed, it can be better focused on the optimal assay design.
The concepts behind bioassay simulation are quite simple, but can become rather complex when several different sources of variation are taken into account.
We focus here on simulation of relative potency assays, but simulation is equally applicable to interpolation type assays (and many others).
Key Takeaways
- By generating realistic data sets based on existing experimental data, bioassay simulation allows researchers to explore various assay designs, dose ranges, plate layouts, and replication strategies, improving efficiency and reducing unnecessary lab experiments.
- Simulation can help determine the optimal number of doses or replicates needed, potentially reducing reagent use and increasing throughput without compromising assay accuracy or precision.
- Simulation can assess the impact of variability sources and test the robustness of an assay under different conditions; subset simulations using real data help identify redundant doses, streamline designs, and validate assay performance even when the true result is not known.
How are the new data simulated?
Relative potency (RP) parallel line analysis and interpolation analysis by either the traditional method or the more efficient BY method all rely on fitting a model (usually linear, or a 4 or 5 parameter logistic) to the data. The mathematical properties of the model are then used in the usual RP or interpolation calculations.
Bioassay simulation is also based on fitting the same initial model, but the aim is to create many more sets of response data. During development, once the behaviour of an assay is at least approximately understood, the data collected to date can be used to build a model of the dose-response relationship.
For simplicity, let’s assume there is only one data set available, with 8 doses, and we wish to create new data sets with 4 replicates at each dose.
The process is as follows:
Step 1
Fit a linear, 4PL or 5PL model to the dataset as usual.

Step 2
Select a set of doses of interest. These can be the original ones or new ones. In this example, we have chosen 10 doses spaced evenly across the range of the data, omitting the 1280 dose as the asymptote is already well defined at a dose of 640.
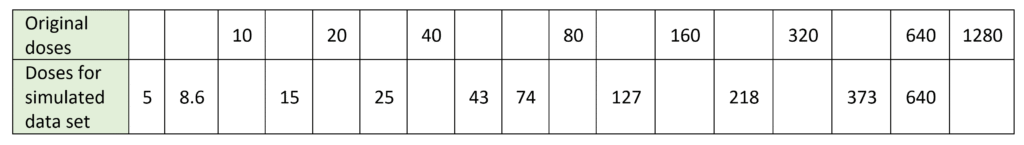
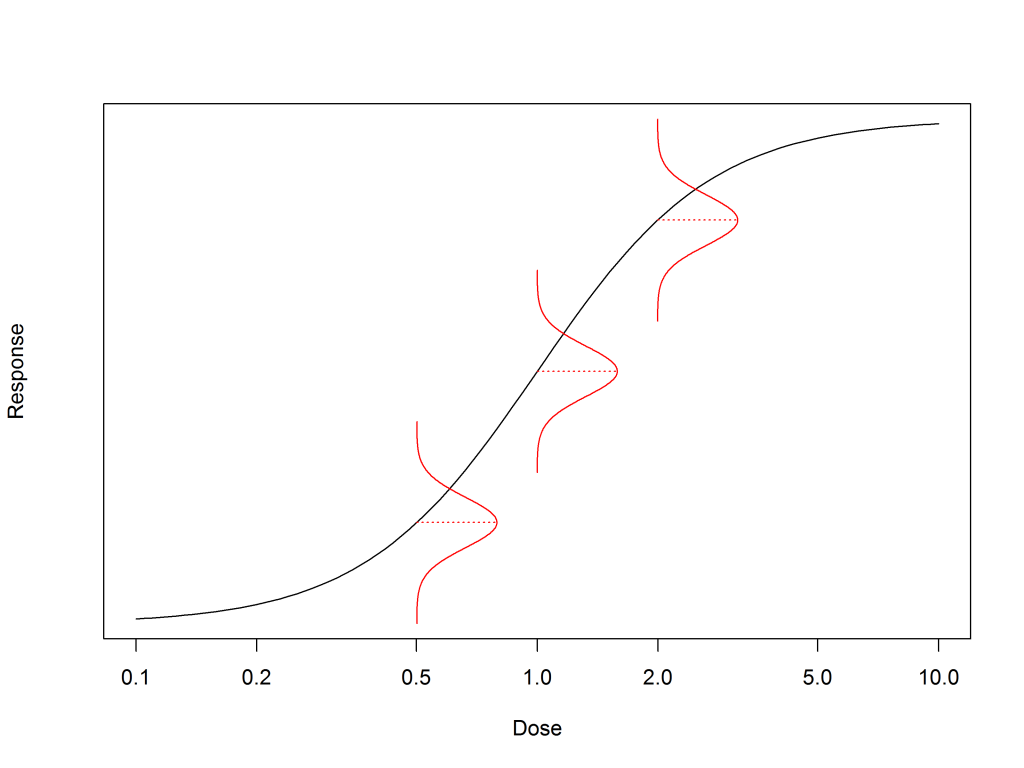
For each dose, read off the response on the model to get the mean response value that is predicted by the model at that dose. Call this md1 for the first dose, d1.

Step 3
To create 4 replicate responses for this dose, we need to know the variability of the real data around the model. As discussed in our blog about homogeneity of variance, the maths of fitting a model in bioassay assumes that the variance of the data is constant across all doses. This variability can be estimated using the root mean square error (RMSE) of all the data from the model.
Step 4
For our 4 replicate responses at each dose we now know the mean and the standard deviation (= RMSE) of the distribution from which they will be sampled.
For the first dose selected, d1, a list of response values can be created that, when taken together, have a normal distribution with a mean of md1 and standard deviation of RMSE. Then 4 of these can be chosen at random as the 4 replicates for this dose. Note that the mean and standard deviation of the 4 replicates chosen will not be md1 and RMSE, but the 4 values have been chosen from a large set of values that have this mean and SD.
This is repeated at each dose of interest and the result is a complete data set.
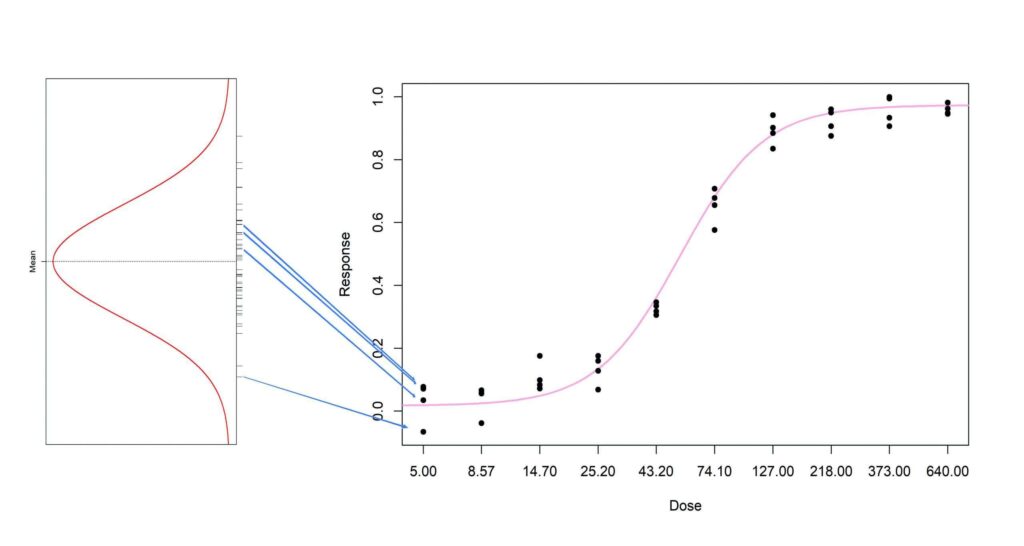
Each time this is done, the data set will be different, but representative of what could occur in real use.
In practice:
- It is not actually necessary to create the big set of simulated response values, but it explains the concepts.
- If there is more than one data set available, then the initial fitted curve will vary a bit between the real data sets – the between plate variability. The curve parameters will have a mean value, and a variability, and this can be added to the simulation. Typically 10–20 data sets will give realistic simulation results.
What type of sample can be simulated?
The process above creates a single new data set. Repeating the process will create another data set; now one can be used as the reference and one as the test sample, this test sample will have a relative potency of 1.0.
For simulating a test (or QC) sample whose RP is 50%, the dose response curve (linear, 4 or 5-parameter logistic), assumed to be parallel to the reference standard curve, simply needs to shift horizontally by log(0.5). That is, the dose associated with a particular mean response is changed to 2 × the original dose.
Using bioassay simulation
Optimising the assay design
Having data sets with a known “true” RP (or concentration) is very useful as it makes it possible to optimise the assay design for throughput, or precision or accuracy (reduced bias) by examining the impact on the calculated RP estimate and precision of, for example:
- Changes in plate layout
- Changes in number of replicates
- Changes in number of plates
- Changes in number of doses / range of doses
Assay sensitivity analysis
This kind of simulation study relies on the same assumptions that are made when the RP and its confidence interval are estimated from the real data. The simulated results are limited by the accuracy of the assumptions.
- The assumptions made can be adjusted to create sensitivity analyses that describe how the assay behaves with changes in the amount and causes of variability.
- Further lab work may be needed to firm up the conclusions.
Simulation using subsets
If a reasonably large amount of data has already been collected (either during development or during routine use), a simpler type of bioassay simulation can sometimes be used to optimise the assay design. This approach involves analysing the data as though only a subset were available, to simulate the case where, for example, fewer dose groups or dilutions had been used.
For example, suppose an assay has been run using 12 doses allowing a 4-parameter logistic model to be used. Then subsets of the real data with perhaps every other dose (6 doses) can be used to calculate the RP, or the concentration in the case of interpolation-type assays. By comparing the results obtained with the full data set and with the subset, this approach can be used to identify redundant doses and increase the throughput of an assay.
The disadvantage of this type of bioassay simulation is that the “true” RP or concentration may not be known (unless the subsets are from the same material e.g. reference vs reference), so the impact of the changes can only be related to the original result, not a known “true” value.
Quantics have recently used this technique for a client to show that 6 similar interpolation type assays can all be changed from 8 doses per sample to 4, thus doubling laboratory throughput, with virtually no impact on the reported concentration estimate or precision.