In the previous parts of our series covering the latest draft version of USP <1033>, we examined the goals of a bioassay validation study and the basics of accuracy and precision. Then, we discussed Probability of Out-of-Specification (Prob(OOS)) and how this can be used to inform the acceptance criteria for accuracy and precision in a validation study. Finally, we covered considerations in a validation study outside of its accuracy and precision, such as dilutional linearity and assay range.
In this final instalment, we’re going to put everything together and walk through an example of the design and analysis of a validation study. This will follow closely with the example provided in appendix A1 of USP <1033> with additional commentary to highlight common confusions and key decision points.
Key Takeaways
- The article shows how a bioassay validation study can be designed end to end, from defining plates, runs, and release procedures through to selecting acceptance criteria and analysing study results.
- Accuracy and intermediate precision are tied back to a risk-based framework, using Prob(OOS) to justify why the chosen limits for relative bias and precision are acceptable.
- The worked example demonstrates how validation conclusions define assay range, showing that acceptable precision and linearity were achieved across the full tested range, but acceptable accuracy was not demonstrated at the highest potency level.
Assays, Runs, and Plates
Before we proceed with the example, however, we must first establish some key terminology. When bioassays are discussed colloquially, the terms “assay”, “plate”, and “run” are often used somewhat interchangeably. In the validation guidance, however, each is used to mean a different – but related – idea, which inevitably becomes confusing. For what follows, we’ll stick to the following definitions:
Plate: A single plate as used in a bioassay which may contain multiple samples. For our purposes, we will assume each plate provides a single relative potency result per sample included on the plate.
Run: A collection of plates that are combined to give a single relative potency result.
Release Procedure: A collection of runs which are combined to give a single relative potency result.
The release procedure format is the specific combination of runs – which might consist of several plates – that is used to generate the reportable result. A very simple format might consist of one run of one plate per run. This would mean the reportable result would be informed by a single plate. A more complicated format might consist of three runs each consisting of two plates. Here, each run would provide a relative potency result which is the combination of the results of two different plates. The assay result would then be a further combination of the results from the three runs. The choice of format is statistically involved and requires investigation of the sources of variability of the assay which is beyond the scope of this blog.
The variability of the final reportable value depends on the release procedure format. In the previous blog, we described this as a measurement variability, but it is also sometimes known simply as the format variability. The measurement variability of the reportable value is defined as
![]()
Where:
![]() is the number of runs in the release assay
is the number of runs in the release assay
![]() is the number of plates per run
is the number of plates per run
![]() is the inter-run variance. This is the variance of the relative potency observed between runs within a release procedure.
is the inter-run variance. This is the variance of the relative potency observed between runs within a release procedure.
![]() is the intra-run variance. This is the variance of the relative potency observed between the plates within a single run.
is the intra-run variance. This is the variance of the relative potency observed between the plates within a single run.
In our example, we will assume the release procedure format (i.e. that which is to be used for routine batch release decisions) is three runs of one plate each. For the validation study itself, we will assume a format of one run consisting of two plates each: this will be referred to as the validation procedure. This reduces the number of plates required in the validation study while still allowing an assessment of the inter- and intra-run variance.
Acceptance Criteria and Analysis Plan
In the previous part of this series, we discussed how the acceptance criteria for a validation study are determined. For accuracy and precision, the choice of acceptance criterion for Relative Bias (RB) and Intermediate Precision (IP) are assessed compared to a desired Prob(OOS). Specifically, we need to determine the Prob(OOS) under the worst allowable performance of the assay: when both the RB and IP are at their acceptance limit.
We also require acceptance criteria for the dilutional linearity and the range of the assay. The table below shows the acceptance criteria we will assume for this example.
| Attribute | Metric | Acceptance Criterion |
|---|---|---|
| Accuracy per level | RB | 90% CI at each level falls within (−11%, +12%) |
| Intermediate precision | IP | 95% UCL for IP at each level (or pooled, if required and appropriate) ≤18% |
| Linearity | Slope | 90% CI on linearity slope within (0.8, 1.2) |
| Range | RB, IP, and slope | Range of levels meeting all acceptance criteria |
For the accuracy and precision of the assay, we have chosen acceptance criteria of ![]() for the relative bias and 18% for the intermediate precision. We can generate the Prob(OOS) for these criteria from the below formula to demonstrate that they are appropriate for this assay:
for the relative bias and 18% for the intermediate precision. We can generate the Prob(OOS) for these criteria from the below formula to demonstrate that they are appropriate for this assay:
![Rendered by QuickLaTeX.com \[Prob(OOS)=\Phi\left(\frac{LSL-\mu_{Process}}{\sqrt{\sigma_{Process}^{2}}}\right)+\left(1-\Phi\left(\frac{USL-\mu_{Process}}{\sqrt{\sigma_{Process}^{2}}}\right)\right)\]](https://www.quantics.co.uk/wp-content/ql-cache/quicklatex.com-a2ffca3190c2708ccc136e058a61de86_l3.png)
Recall that we define the “true” relative potency as the mean of the product distribution, ![]() . However, we can only ever observe estimates of the potency using the bioassay. The mean of the distribution of the observed results is then
. However, we can only ever observe estimates of the potency using the bioassay. The mean of the distribution of the observed results is then ![]() . The difference between
. The difference between ![]() and
and ![]() is the relative bias of the assay. The maximum bias allowable for the assay is 12%, so, if we assume that
is the relative bias of the assay. The maximum bias allowable for the assay is 12%, so, if we assume that ![]() , we find that
, we find that ![]() for our calculation.
for our calculation.
For the precision, we defined the variance of the process distribution as ![]() . For the release assay format of
. For the release assay format of ![]() ,
, ![]() , we can express the measurement variability in terms of the intermediate precision as
, we can express the measurement variability in terms of the intermediate precision as
![]()
Inputting our IP criterion of 18%, we find that, in the worst allowable case, ![]() . We cannot directly calculate
. We cannot directly calculate ![]() from the properties of the process distribution. Instead, we can make an assumption about the relative contributions of the product and measurement variability to the process variability. A typical breakdown for a bioassay is that 20% of the variability is inherent to the product while 80% comes from the measurement. By substituting
from the properties of the process distribution. Instead, we can make an assumption about the relative contributions of the product and measurement variability to the process variability. A typical breakdown for a bioassay is that 20% of the variability is inherent to the product while 80% comes from the measurement. By substituting ![]() into
into ![]() , we find that
, we find that
![]()
Now, let’s assume the specification limits for the relative potency of our product are (70%, 143%). We now have all the ingredients we need to calculate the Prob(OOS):
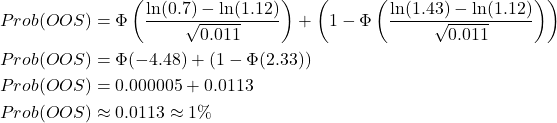
That tells us that acceptance criteria of 12% for the RB and 18% for the IP represent good choices for this assay, since the worst-case Prob(OOS) using these criteria is expected to be about 1%.
In order, then, to properly assess our assay in the validation study, we must complete the following analyses:
- Plot and trend measured relative potency against target relative potency to assess dilutional linearity
- Calculate the %RB at each potency level, and check its 90% CI falls within the acceptance bounds of (-11%, 12%)
- Calculate the IP at each potency level, and check the upper confidence limit on its 95% CI falls below 18%. If required and appropriate, this may be evaluated as an estimate pooled across potency levels.
Study Design and Sample Size
A validation study is designed to demonstrate the performance of the assay in the conditions it will encounter in routine use. This includes testing under the allowed range of factors which may affect the results of the assay, such as different analysts or reagent lots. The goal of the study is not to determine these allowed ranges or test the effect of these factors – this should have been established during the design of the assay using design of experiments and robustness studies respectively. Instead, we aim to establish the performance of the assay in the presence of allowed variation.
The validation study should be conducted over a range of representative potency levels. These should minimally cover the specification range of the product, but levels outside the specification are often also included to assess assay performance for out-of-specification lots and stability samples. The choice of potency levels should also be appropriate to establish dilutional linearity, meaning at least three, and ideally five, potency levels should be included. For our example, we have considered a product specification range of ![]() . A good choice of five potency levels might be, therefore,
. A good choice of five potency levels might be, therefore, ![]() . This covers the specification range of the assay with three levels, and includes a level each outside the USL and the LSL.
. This covers the specification range of the assay with three levels, and includes a level each outside the USL and the LSL.
To calculate the sample size, we must find the smallest number of assays required to give a high likelihood of drawing the correct conclusion based on the data collected. We’ve covered sample size calculations in great detail in the context of clinical trials, so we won’t go into detail about the statistical fundamentals here. Appendix A2 of draft USP <1033> quotes the formula for calculating the required sample size for conformance on a relative bias criterion as
![Rendered by QuickLaTeX.com \[n \geq \frac{\left(t_{\alpha,df}+t_{\frac{\beta}{2},df}\right)^{2} \times \hat{\sigma}_{IP}^{2}}{\theta^{2}}\]](https://www.quantics.co.uk/wp-content/ql-cache/quicklatex.com-b538878bc995b7da4222621262a87990_l3.png)
Where:
![]() gives the critical value from the t-distribution with upper tail probability
gives the critical value from the t-distribution with upper tail probability ![]() and degrees of freedom
and degrees of freedom ![]() . In this case,
. In this case, ![]()
![]() is the estimated log scale variance (from historical data, for example).
is the estimated log scale variance (from historical data, for example).
![]() is the acceptance criteria for the %RB on the log scale.
is the acceptance criteria for the %RB on the log scale.
![]() is the one-sided confidence level for the confidence interval on the relative bias. Typically,
is the one-sided confidence level for the confidence interval on the relative bias. Typically, ![]() .
.
![]() is the two-sided risk of failure when the RB is zero. Here,
is the two-sided risk of failure when the RB is zero. Here, ![]() .
.
This formula is implicit – to define the sample size ![]() , we must know the degrees of freedom
, we must know the degrees of freedom ![]() . That means we must use an iterative trial-and-error method to find an
. That means we must use an iterative trial-and-error method to find an ![]() which satisfies our requirements. Specifically, we iterate until the sample size we input when calculating the degrees of freedom (
which satisfies our requirements. Specifically, we iterate until the sample size we input when calculating the degrees of freedom (![]() ) is greater than the output from the formula (
) is greater than the output from the formula (![]() ). The table below shows an example of this iterative calculation based on the below inputs:
). The table below shows an example of this iterative calculation based on the below inputs:
![]() (based on an assumed IP of 7%)
(based on an assumed IP of 7%)
![]() (based on a %RB criterion of 12%)
(based on a %RB criterion of 12%)
![]()
![]()
![]()
| Inputs | Try n | Calculated | ||||||
|---|---|---|---|---|---|---|---|---|
| |
||||||||
| 0.068 | 0.11 | 0.05 | 0.025 | 4 | 3 | 2.353 | 3.182 | 10.9 |
| 5 | 4 | 2.132 | 2.776 | 8.6 | ||||
| 6 | 5 | 2.015 | 2.571 | 7.5 | ||||
| 7 | 6 | 1.943 | 2.447 | 6.9 | ||||
| 8 | 7 | 1.895 | 2.365 | 6.5 | ||||
We see that ![]() , as calculated using the formula, is greater than
, as calculated using the formula, is greater than ![]() for
for ![]() . This flips once
. This flips once ![]() , which means that a sample size of
, which means that a sample size of ![]() is acceptable. For our example, we will choose to use a sample size of
is acceptable. For our example, we will choose to use a sample size of ![]() as this allows for a balanced study design and means the analysis is more robust to inaccuracies in the assumptions – if the intermediate precision was greater than expected, for example.
as this allows for a balanced study design and means the analysis is more robust to inaccuracies in the assumptions – if the intermediate precision was greater than expected, for example.
Note that, since we used the intermediate precision to calculate the sample size, it strictly applies to the number of plates. However, we have chosen to apply the sample size to the number of procedures as this allows us to evaluate within-procedure variance and does not lead to an extreme use of resources.
The validation procedure format consists of one run of two plates. On each of these plates, we include the reference and a test sample from each of the chosen potency levels. Our required sample size is , meaning we must perform the procedure eight times in the validation study.
The key factors which were identified in a DOE study were the influence of the reagent lots and the analyst, so we must ensure our study design accounts for both of these. The diagram below outlines a study design which might be appropriate in such a scenario.
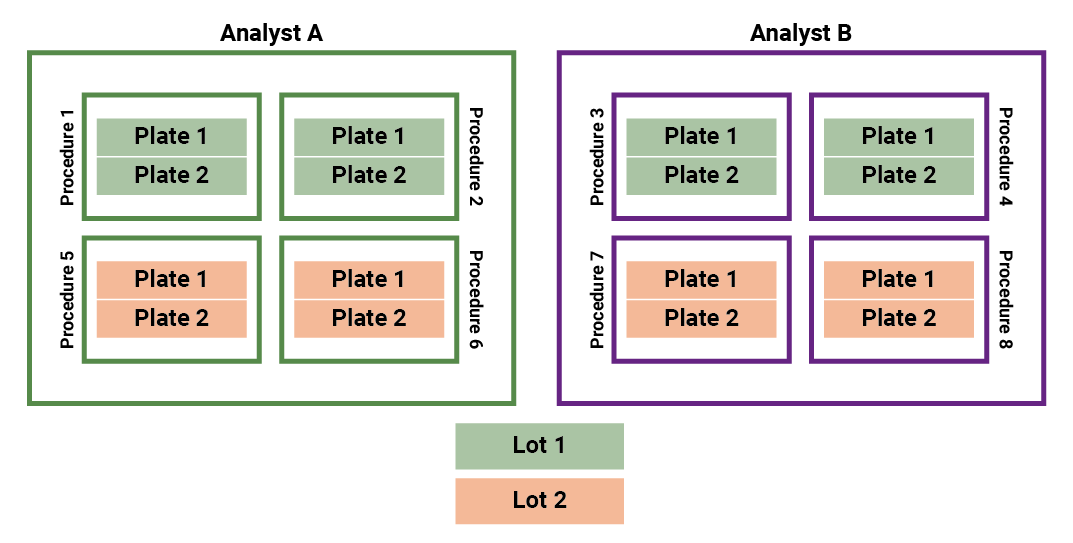
The assays are conducted by one of two analysts, Analyst A or Analyst B, and use one reagent Lot 1 or Lot 2. Each analyst conducts four procedures, two using Lot 1 and two using Lot 2. This ensures that the design is balanced across the two key factors which have been identified for this assay. Following the designated format, each procedure consists of one run of two plates. That means a total of sixteen plates are run across the full study. Each plate contains a sample of each target potency, meaning the study design achieves the required sample size of eight procedures once the relative potency results from each plate are combined into a single reportable value.
The Validation Analysis
With the study designed and the analysis laid out, all that is left is to collect the data and perform the calculations. Here, we’ll give an overview of the assessment of accuracy and precision for a single potency level, before zooming out to examine dilutional linearity and the process of forming conclusions across the full range of potency tested in the study.
The table below shows an example of data which might be collected for the 50% potency level. Recall that the assay format for the validation study consists of one run of two plates. That means that, from the eight assays performed for this potency level, we have sixteen total measurements of the relative potency. We will use this data to evaluate the accuracy and precision of the assay at the 50% potency level.
| Procedure | Analyst | Lot | RP 1 | RP 2 | Geomean(RP) |
|---|---|---|---|---|---|
| 1 | A | 1 | 0.5215 | 0.5026 | 0.511963 |
| 2 | A | 1 | 0.4532 | 0.4497 | 0.451447 |
| 3 | A | 2 | 0.5667 | 0.5581 | 0.562384 |
| 4 | A | 2 | 0.5054 | 0.535 | 0.519989 |
| 5 | B | 1 | 0.5222 | 0.5017 | 0.511847 |
| 6 | B | 1 | 0.5179 | 0.5077 | 0.512775 |
| 7 | B | 2 | 0.5314 | 0.5411 | 0.536228 |
| 8 | B | 2 | 0.5112 | 0.5488 | 0.529666 |
Accuracy
The accuracy of the assay is evaluated based on the relative bias, which is given by:
![]()
Where ![]() is the measured relative potency and
is the measured relative potency and ![]() is the target relative potency. In our case, the measured relative potency is the geomean of the averaged relative potency measurements from assay:
is the target relative potency. In our case, the measured relative potency is the geomean of the averaged relative potency measurements from assay: ![]() . The target relative potency is 50%, so:
. The target relative potency is 50%, so:
![]()
Our acceptance criterion is that the 90% CI on the relative bias must lie entirely within the acceptance bounds. We won’t go into detail on the calculation of the confidence intervals here – this can be done using statistical software. After performing this calculation, we find the result:
![]()
That means the 90% CI for the relative bias falls entirely within the acceptance bounds of , meaning we have demonstrated acceptable accuracy at the 50% potency level.
Precision
The precision of our assay is evaluated using an acceptance criterion on the intermediate precision, which can be calculated as:
![]()
A Variance Components Analysis (VCA) can be used to calculate ![]() and
and ![]() . This calculation is beyond the scope of this blog – we’ll cover VCA in a future blog. For this data, we find that
. This calculation is beyond the scope of this blog – we’ll cover VCA in a future blog. For this data, we find that ![]() and
and ![]() . That gives us an IP of
. That gives us an IP of
![]()
Accuracy and Precision Across Levels
These calculations are performed for the data collected at each target potency level. An example of results for the five chosen levels is shown in the table below.
| Target Potency (%) | Relative Bias 90% CI (%) | Intermediate Precision (%) |
|---|---|---|
| 50 | (-1.02, 7.67) | 6.8% |
| 71 | (-3.42, 3.67) | 7.3% |
| 100 | (0.06, 10.12) | 8.5% |
| 1.41 | (-1.04, 7.03) | 6.3% |
| 2 | (5.31, 14.32) | 7.2% |
Notice that, while we have calculated a confidence interval for the relative bias at each level, a similar procedure has not been performed for the intermediate precision despite the acceptance criterion being based on the upper 95% confidence limit. This is because the quantity of data at each level is limited relative to the overall study design. Instead, for data which is sufficiently similar (i.e., if the maximum variance component is less than ten times the minimum), we can calculate a pooled estimate of the intermediate precision. This is used to assess all five potency levels against the acceptance criterion using a confidence interval for the pooled estimate. In this case, the upper confidence limit on the pooled intermediate precision is 11.8%. Since this is within the acceptance limit of 18%, we can conclude that the precision is acceptable for all potency levels.
The same, however, cannot be said for the accuracy of the assay. This becomes apparent when the 90% confidence intervals are plotted for each potency level, as in the plot below.
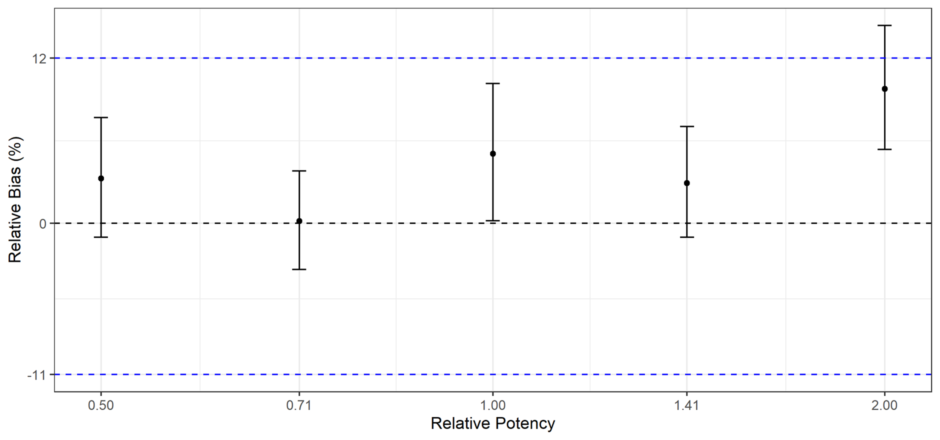
The blue lines represent the acceptance limits. We can clearly see that, while the 90% CIs for the lower four potency levels fall within the acceptance bounds, that for the 200% level does not. That means the assay has not demonstrated acceptable accuracy for that potency level.
The final component required to make a full assessment on the validity of the assay is to examine the dilutional linearity. To do this, we plot the measured relative potencies against the target relative potency at each of the potency levels, with both on the log scale, and fit a linear regression model. This is shown in the plot below. Each point in the plot represents the geometric mean of the relative potency returned by each procedure, meaning there are four points per analyst per potency level.
The acceptance criterion for the dilutional linearity study is based on the slope of the linear relationship between the measured and target relative potency. So, fitting a linear regression model, we find that the relationship is best described by:
![]()
The fitted slope was determined as 1.04 with a 90% CI of (1.02, 1.07). This falls within the acceptance bounds of (0.8, 1.2), meaning we can conclude that the assay is acceptably linear over its full range.
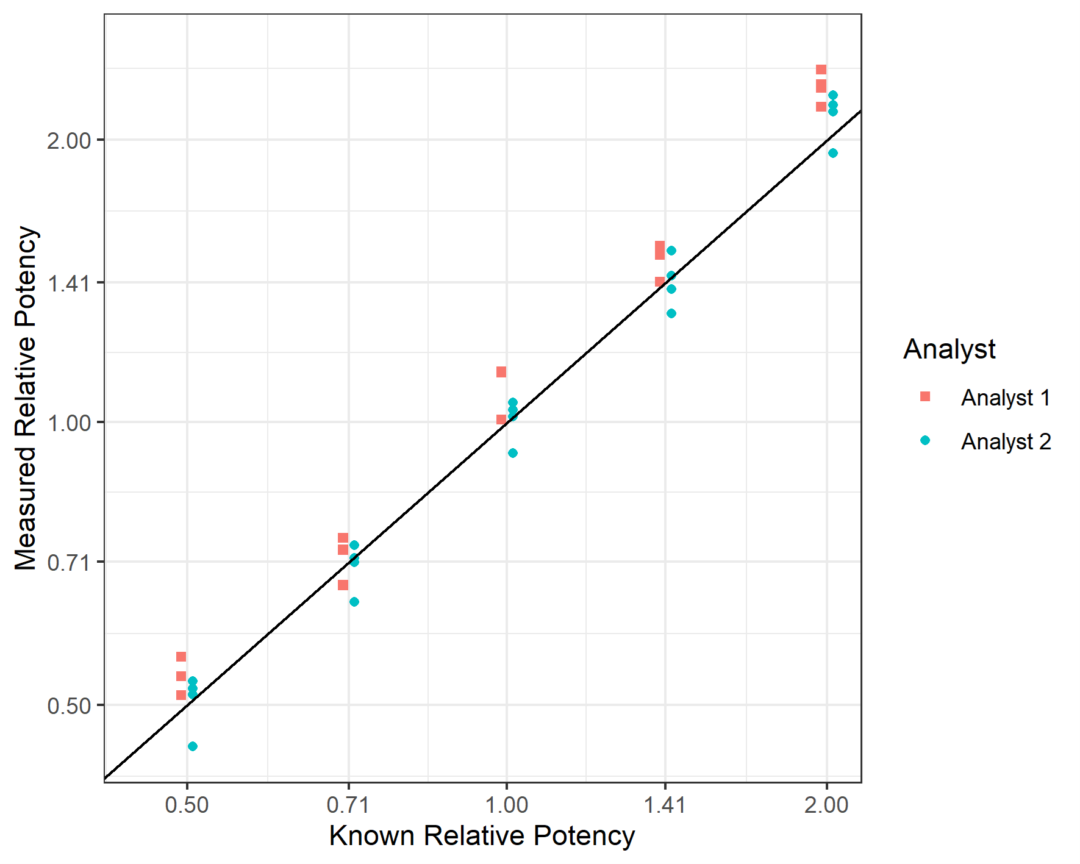
Our overall conclusion, therefore, is that the assay shows acceptable accuracy, precision, and linearity at target potencies between 50% and 141%. At a target potency of 200%, the accuracy of the assay was not shown to be acceptable. This means we state that the assay has been validated for an assay range of 50%-141%.
To Jump a Hurdle
Bioassay validation is the culmination of development: the final test which an assay must pass in order to serve its purpose in the testing of therapeutics. It is, understandably, a daunting process, with a wide range of considerations required to ensure the assay is tested rigorously before routine use. With the right planning – and early statistical involvement – however, the complexity of validating a bioassay can be minimised, ensuring that the time, effort, and resources utilised in developing and testing a bioassay are not wasted.






Comments are closed.