So far in our series examining approaches to outlier management in bioassay, we’ve discussed the various perspectives on outliers held by stakeholders in the bioassay lifecycle and the problems the presence of outliers can pose for an assay system. We then used a simulation to evaluate the performance of a couple of methods for outlier detection – Grubbs’ Test and Studentised Residuals. We still have, however, a question left unanswered: what happens if our outlier detection process flags legitimate points as outliers?
In an ideal world, we would have evidence beyond a statistical test – such as knowledge of an error in the lab – which allows us to differentiate outlying points from the inherent variability of the assay. As we’ve previously established, however, this is often not possible, meaning we are relying on statistical methodologies to determine whether a point is an outlier. These tests can either be correct – identifying an outlying point – or can fail in one of two ways. They can miss outliers, incorrectly identifying an outlier as a part of the inherent variability of the assay, or they can incorrectly identify a legitimate point as an outlier.
Key Takeaways
-
Outlier misidentification is inevitable when using statistical tests due to Type I error; tuning significance levels trades sensitivity for specificity, so some legitimate observations will be flagged regardless.
-
Removing legitimate points reduces accuracy and can distort inference: in a simulation with no true outliers added, datasets with removals showed higher AAD (worse accuracy) and artificially improved PF (apparent precision), which also increased erroneous parallelism test failures.
-
Mitigation is possible without blanket deletion: use outlier flagging with contextual checks, consider robust/ accommodation methods (e.g., robust regression), and control precision via acceptable confidence-interval widths rather than defaulting to point removal.
We know that missing outliers can prove problematic: we’ve demonstrated that outliers decrease the accuracy and precision of relative potency measurements. But does the misidentification and removal of legitimate points cause similar problems? We can certainly see why unnecessary removal of data would be an issue from a scientific perspective. By removing misidentified points, we are losing valuable information about our sample. Much of the bioassay design process centres around including as many observations as efficiently as possible, so to throw one or more out for no good reason seems to fly in the face of the goals of bioassay analysis.
Clearly, then, we want to avoid misidentifying and removing legitimate observations as outliers. Unfortunately, implementing any form of statistical outlier detection method will result in a small number of observations being misidentified. This is due to the Type I error rate of the statistical test. A Type I error is a false positive: the statistical test concludes there is evidence for the outcome we are trying to detect (in this case, whether an observation is outlying) due to chance variability of the data.
For example, imagine you were testing a coin to see whether it was fair. You assume it’s fair, and are looking for evidence that its biased based on the longest run of a single result being returned. You flip the coin five times, and it returns five heads in a row, which is enough evidence for you to conclude that the coin is biased. But hold on! A fair coin would be expected to return five heads in a row (or five tails in a row) once out every 32 5-flip sequences (![]() ). That means there’s a one in 32 (
). That means there’s a one in 32 (![]() ) chance that a fair coin would be misidentified as biased purely by chance.
) chance that a fair coin would be misidentified as biased purely by chance.
One could decrease the chance of misidentification by increasing the length of the run required before a coin was considered biased – increasing the run length to 10 would decrease the chance of misidentification to one in 1024 (![]() ). This is known as decreasing the significance level of the test. But there’s a trade-off here: the lower the significance level, the more evidence is required before the test returns a positive result. That means there’s a greater chance of a false negative (a Type II error). Here, that would be failing to identify a truly biased coin as such and concluding that it was fair.
). This is known as decreasing the significance level of the test. But there’s a trade-off here: the lower the significance level, the more evidence is required before the test returns a positive result. That means there’s a greater chance of a false negative (a Type II error). Here, that would be failing to identify a truly biased coin as such and concluding that it was fair.
This trade-off exists for any statistical test – including outlier tests – meaning if we are to have a reasonable chance of detecting truly outlying observations, then we must accept that there will be some legitimate observations that are misidentified as outliers. Of course, we always have the choice to set the significance level to our liking. A test with a higher significance level will be more sensitive – less likely to miss a true outlier at the cost of a greater misidentification rate – while a test with a lower significance level will be more specific – less likely to misidentify legitimate observations as outlying, but more likely to miss a true outlier.
So, given the misidentification of legitimate observations as outliers is an inevitable consequence of employing a statistical outlier test, what is the consequence of outlier misidentification for the accuracy and precision of a relative potency measurement? We’ve run – you guessed it – a simulation to demonstrate the effects of removing legitimate observations on bioassay results.
The setup for this simulation was much the same as that for the simulations run in the previous two parts of this series. The difference here is that, unlike previously, no outliers were added to the simulated data. All observations in each of the six dose groups were drawn from the same distribution with ![]() . According to the definition of an outlier, therefore, none of the observations would be considered outlying as they are all drawn from the same statistical population.
. According to the definition of an outlier, therefore, none of the observations would be considered outlying as they are all drawn from the same statistical population.
1000 such datasets were generated, and analysed in QuBAS using two methods, one using Grubbs’ test and one using Studentised Residuals. As before, Grubbs’ test used a threshold (i.e. a significance level) of ![]() for outlier detection, while the Studentised Residuals approach used a threshold of
for outlier detection, while the Studentised Residuals approach used a threshold of ![]() . In both cases, the analysis method also included an F Test for parallelism with a significance level of . Once analysed, each dataset was sorted into categories based on how many “outliers” were detected. These were:
. In both cases, the analysis method also included an F Test for parallelism with a significance level of . Once analysed, each dataset was sorted into categories based on how many “outliers” were detected. These were:
![]() : No observations were identified as outliers
: No observations were identified as outliers
![]() : A single observation was identified as an outlier and removed
: A single observation was identified as an outlier and removed
![]() : Multiple observations were identified as outliers and removed
: Multiple observations were identified as outliers and removed
The accuracy of the relative potency measurements was once again assessed using the Average Absolute Deviation (AAD), while the precision of individual relative potency measurements was encapsulated using the Precision Factor (PF). The percentage of datasets which fell into each detection category and the key metrics for each of those categories are shown in the plot and table below respectively.
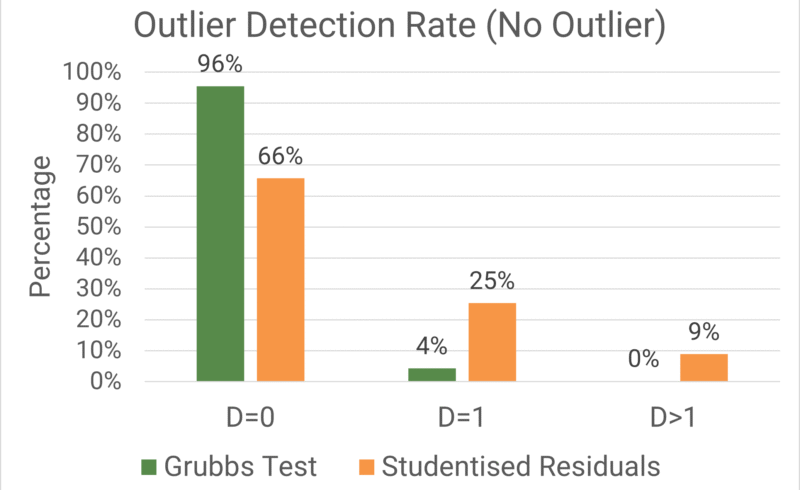
As we might expect, ![]() was the most frequent category for both Grubbs’ test and Studentised Residuals. There were no true outliers present in the data, so it would be a big surprise if
was the most frequent category for both Grubbs’ test and Studentised Residuals. There were no true outliers present in the data, so it would be a big surprise if ![]() did not contain the majority of datasets for both methods. There is, however, a noticeable difference in how large a majority
did not contain the majority of datasets for both methods. There is, however, a noticeable difference in how large a majority ![]() contained between the methods. For Grubbs’ test, more than 95% of datasets tested fell into the
contained between the methods. For Grubbs’ test, more than 95% of datasets tested fell into the ![]() category, while this was the case for approximately 66% of datasets for Studentised Residuals.
category, while this was the case for approximately 66% of datasets for Studentised Residuals.
We set the significance level – the Type I error rate – of Grubbs’ test to be 5%, so we would expect that 5% of datasets would return a false positive. In this case, this false positive is the misidentification of an outlier. For Studentised Residuals, by contrast, the threshold of ![]() meant 34% of datasets returned a false positive. As such, Studentised Residuals is a more sensitive outlier test in this configuration, while Grubbs’ test is more specific.
meant 34% of datasets returned a false positive. As such, Studentised Residuals is a more sensitive outlier test in this configuration, while Grubbs’ test is more specific.
Since so few datasets showed ![]() for Grubbs’ test, we’ll focus on the results from Studentised Residuals, which has illustrative sample sizes for both
for Grubbs’ test, we’ll focus on the results from Studentised Residuals, which has illustrative sample sizes for both ![]() and
and ![]() . The AAD for
. The AAD for ![]() (11.7%) was lowest among the categories, with an increase in AAD observed for both
(11.7%) was lowest among the categories, with an increase in AAD observed for both ![]() (14.4%) and
(14.4%) and ![]() (15.4%). This tells us that the measured relative potency results for
(15.4%). This tells us that the measured relative potency results for ![]() were, on average, further away from the true relative potency of 100% than those for
were, on average, further away from the true relative potency of 100% than those for ![]() . In essence, the misidentification of outliers decreases the accuracy of the assay system.
. In essence, the misidentification of outliers decreases the accuracy of the assay system.
By contrast, the precision of the relative potency measurement increases when observations are misidentified as outliers. The precision factor for ![]() (1.79) was greater than that for
(1.79) was greater than that for ![]() (1.72) and
(1.72) and ![]() (1.67). This might seem surprising – surely more observations lead to a more precise measurement? The clue lies in which observations are being removed: the outlier test preferentially removes the observations which are furthest from the model. These are the observations which add the most variability, so the precision of the relative potency measurement is artificially increased. This is not a good outcome: it misrepresents the true confidence intervals of the relative potency measurement, which can lead to incorrect conclusions being reached in batch testing, say.
(1.67). This might seem surprising – surely more observations lead to a more precise measurement? The clue lies in which observations are being removed: the outlier test preferentially removes the observations which are furthest from the model. These are the observations which add the most variability, so the precision of the relative potency measurement is artificially increased. This is not a good outcome: it misrepresents the true confidence intervals of the relative potency measurement, which can lead to incorrect conclusions being reached in batch testing, say.
We see another consequence of the artificial variability reduction in the behaviour of the F Test for parallelism. Just as with the statistical outlier tests, the F Test has a Type I error rate: here we set this to be ![]() . Since the data were simulated to be truly parallel, we would expect 1% of the datasets to fail the parallelism test by chance. And indeed, this is what we observe for the
. Since the data were simulated to be truly parallel, we would expect 1% of the datasets to fail the parallelism test by chance. And indeed, this is what we observe for the ![]() category, for which 98.6% of datasets passed the parallelism test. For
category, for which 98.6% of datasets passed the parallelism test. For ![]() and
and ![]() , however, this reduced to 94.9% and 86.5respectively. This was a result of erroneous F Test failures due to the artificially low variability of the data when misidentified observations were removed.
, however, this reduced to 94.9% and 86.5respectively. This was a result of erroneous F Test failures due to the artificially low variability of the data when misidentified observations were removed.
So, we can conclude that the misidentification and removal of legitimate observations as outliers causes inaccuracy in relative potency results. However, if misidentification is an inevitable consequence of using statistical outlier tests, what can we do to minimise the effect they have on our results?
One approach could be to add checks between a potential outlier being detected and the decision to remove the observation from the analysis. Known as outlier flagging, this process allows for information beyond the output of a statistical test to be incorporated into decision-making. For example, one could use historical information about the variability of the assay to determine whether a potential outlier is likely to have been flagged by chance or whether it is a genuine outlier. Flagging an outlier for further investigation before removal also provides an opportunity to examine whether that observation can be explained by a lab error, deviation from procedure, or some other atypical event. This information would give greater support to the decision to remove the observation. There will always be some degree of subjectivity associated with such decisions, but, provided a rigorous framework exists to guide the process, outlier flagging can mitigate some of the negative effects of outlier misidentification.
What if we chose not to remove outliers altogether? Statistical analysis approaches exist which can accommodate the presence of outliers while mitigating the bias which would otherwise be introduced. A very simple example of this is using the median instead of the mean when summarising a set of a data. Imagine five schoolchildren took a spelling test and returned the following scores out of 20: {17, 18, 16, 18, 7}. The mean score of the group is 15.2: 4/5 of the students – 80%! – scored higher than this on the test, so it’s not a particularly good representation of the performance of the group. The score of 7, which we could consider an outlier, has had an undue influence on the mean score in the group. The median score, however, is 17. This is a far better representation of the performance of the students, largely due to the mitigation of the effect of the outlying result.
How does this apply to a bioassay? In this context, outlier accommodation techniques can be a little more complex, so we recommend discussing these with a statistician. For single point outliers, which have been the main focus of our investigations, techniques such as robust regression provide model fitting approaches which are designed not to be affected by the presence of outliers. While computationally expensive, this allows analysis to proceed without the need for outlier detection and removal.
Perhaps the simplest approach is to make use of the information you’re likely to be collecting anyway: confidence intervals. As we have seen, outliers affect the precision of a reportable value, meaning the width of said confidence interval will reflect whether there were outliers present in the data. By setting limits on the confidence intervals of estimated values, the precision can be controlled and, by proxy, so can the impact of outliers. If the precision is not sufficiently affected that the confidence interval is unacceptably wide, however, then one could make the case that there is no reason to be concerned. It is impossible for an outlier to increase the precision of an estimate, so we can be confident that if the precision of our result is acceptable, then this is in spite of any outliers rather than because of them.
As with all the aspects of outlier management we’ve investigated in this series, outlier misidentification is a complex issue with several consequences and approaches for mitigation. The removal of legitimate observations can cause bias in reportable values, while also artificially decreasing the variability of the data due to the preferential removal of the observations with the most variance. This can not only lead to incorrect conclusions being drawn about the assay results, but can also cause problems for some methods for suitability testing. All statistical outlier detection methods, however, will inevitably result in some outlier misidentification. Outlier flagging is an approach which can mitigate the effects of misidentification, while alternative approaches can be employed to remove the need for outlier detection altogether.






Comments are closed.