When is Pure Error for Bioassay an Error?
Software systems for calculating relative potencies and other bioanalytical results often provide a choice of variance estimation method, usually: Residuals, Lack of fit Error and Pure Error. This defines how the confidence limits on the reportable result are calculated. So which should you choose?
The answer is simple. If the reportable result is calculated in a process that involves fitting a model to the data, e.g. linear, 4pl, 5pl, then you should always choose Residuals. Unfortunately, Pure Error sounds good and many scientists choose this. It may also be the default setting in some software packages, but choosing this will result in confidence intervals that are falsely narrow.
This is really an easy choice once explained. Consider a simple linear model fit to a reference standard and sample which have 4 replicates each. This is shown in Figure 1.
Key Takeaways
- Variance estimation methods differ in what aspects of variability they measure—Residuals, Lack of Fit, and Pure Error are not interchangeable.
- Residuals should always be selected when the reportable value is derived from a fitted model, as it reflects both replicate variability and model fit.
- Choosing Pure Error or Lack of Fit underestimates true variability, producing confidence intervals that are misleadingly narrow.
- Residuals provide the most realistic assessment of assay variability and therefore the most reliable confidence limits.
Also in Figure 1, we have focused in on one of the dose groups to help show how Pure Error and Lack of Fit are calculated.
Pure Error in light blue only measures the variability of the replicates from the mean of the replicates, at each dose, but the fit to the model is ignored.
Residuals combines these errors and takes account of both:
- the variability of the replicates from the mean of the replicates
- and
- the lack of fit of the mean of the replicates to the fitted model.
Lack of fit Error in dark blue only measures the lack of fit of the mean of the replicates to the fitted model, ignoring the replicate variability.
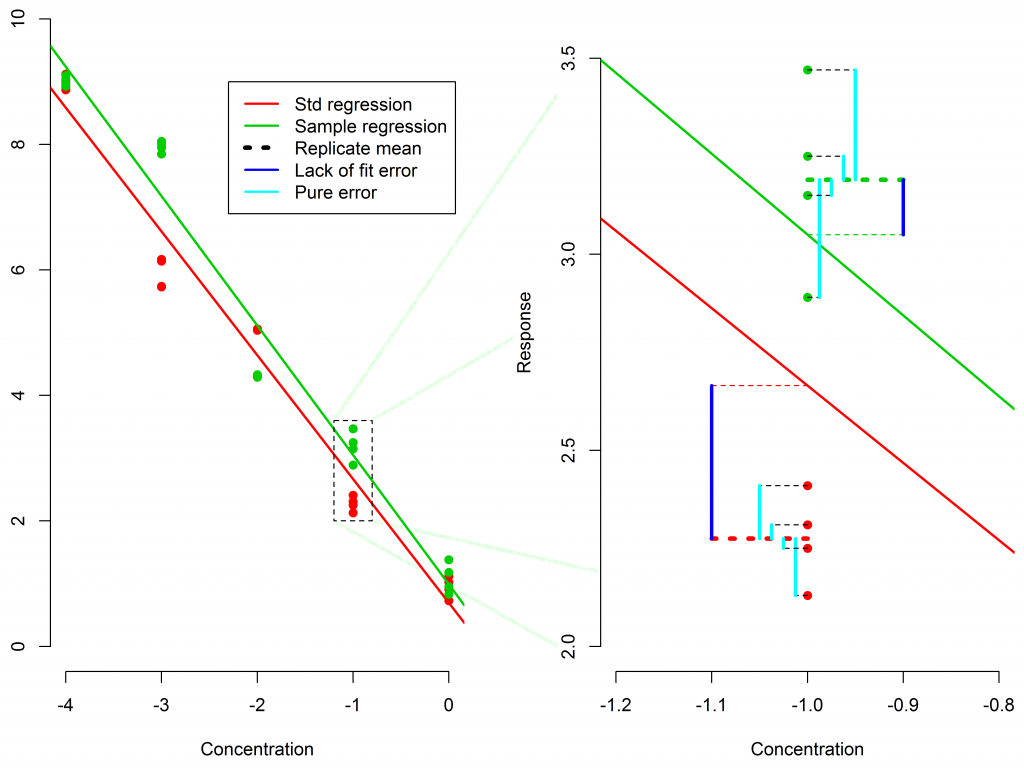
So clearly, using Pure Error or Lack of fit Error will underestimate the actual variability of the data with respect to the fitted model used to calculate the reportable value. The confidence intervals for the reportable value are calculated based on this variability, so the confidence limits will be falsely narrow.
Always use Residual error.
Simple!