Statistical Model Formulae: The BEBPA Sessions
Myself, and my colleague Brooke, attended the BEBPA US virtual meeting last week and reported back on the presentation and round table discussions on relative potency software, including some questions about the use of different types of statistical model.
The key driver for these sessions was the experience that many have had of different software and different versions of the same software giving different results, and the problems this causes with RP values, and system and sample suitability criteria when transferring assays to different labs.
The discussions focussed around the concept of bridging between software using test data sets. However, the reasons for the differences seen were not clearly explained and are in fact just mathematical. No “bridging” exercise, in the sense of testing a few data sets on both systems as suggested, is required. In fact, doing so may simply provide a false sense of security.
In light of these discussions, we worked with the team here at Quantics to put together a series of 3 short blogs. These are structured to look at the key issues so that software users can understand what is required to “bridge” software mathematically, and why there may still be differences.
Key Takeaways
- Although various regulatory documents (like USP and PhEur) present 4PL formulas differently, they are just different parameterisations of the same mathematical model. No experimental “bridging” is needed between software systems — a simple algebraic conversion ensures identical results.
- Discrepancies between relative potency results across software platforms are usually due to differences in fitting algorithms rather than differences in the mathematical formula. Understanding this distinction is key to proper assay validation and software bridging.
- With proper mathematical conversion between parameterisations, assay validation remains intact when switching or comparing software. This saves significant time and avoids unnecessary re-validation, provided the curve-fitting algorithms are robust and well understood.
Blog 1. Equations and algorithms.
During the presentation there was a degree of confusion between the terms equation or formula and algorithms, but they are different and the difference is important to aid understanding. An equation or formula is a mathematical relationship or rule expressed in symbols. An algorithm is a finite sequence of well-defined instructions, typically used to solve a class of specific problems or to perform a computation.
The presenter stressed that there were several different formulae used for a 4PL and that these give rise to different results. We were told that:
- It was important that both sets of software use the same one
- The formulae chosen was the one that the regulators support.
This is incorrect.
It is true that the USP, PhEur and other mathematical references give examples of apparently different equations for the 4PL, but the key point is that they are all mathematically identical, and will therefore always give identical results. The formulae differ in how the parameters in the equation are defined. This is called parameterisation and the regulators will accept any parameterisation as it makes no difference to the results (RP and confidence intervals).
Brownies
It is a bit like a recipe for your favourite brownies. In the formula (ingredients and temperatures) the quantities can be expressed in ounces or grams, the temperature in degrees centigrade or Fahrenheit. Once you know this, it is simple to “translate” from one to the other. You don’t have to do a bridging exercise making the brownies in both ways (unless you really like brownies.)
| Metric parameterisation | US parameterisation | |
|---|---|---|
| Oven temperature | 180°C | 347°F |
| 80% chocolate | 245g | 8.642oz |
| Butter | 200g | 7.055oz |
| Easter sugar | 125g | 4.41oz |
| Dark soft brown sugar | 125g | 4.41oz |
| Salt | 0.4 (1 pinch) | 0.014oz |
| Plain flour | 115g | 4.051oz |
| Eggs | 4 | 4 |
| Vanilla extract | 2 tsp | 0.17oz |
| Pecan nuts | To taste | To taste |
Linear model
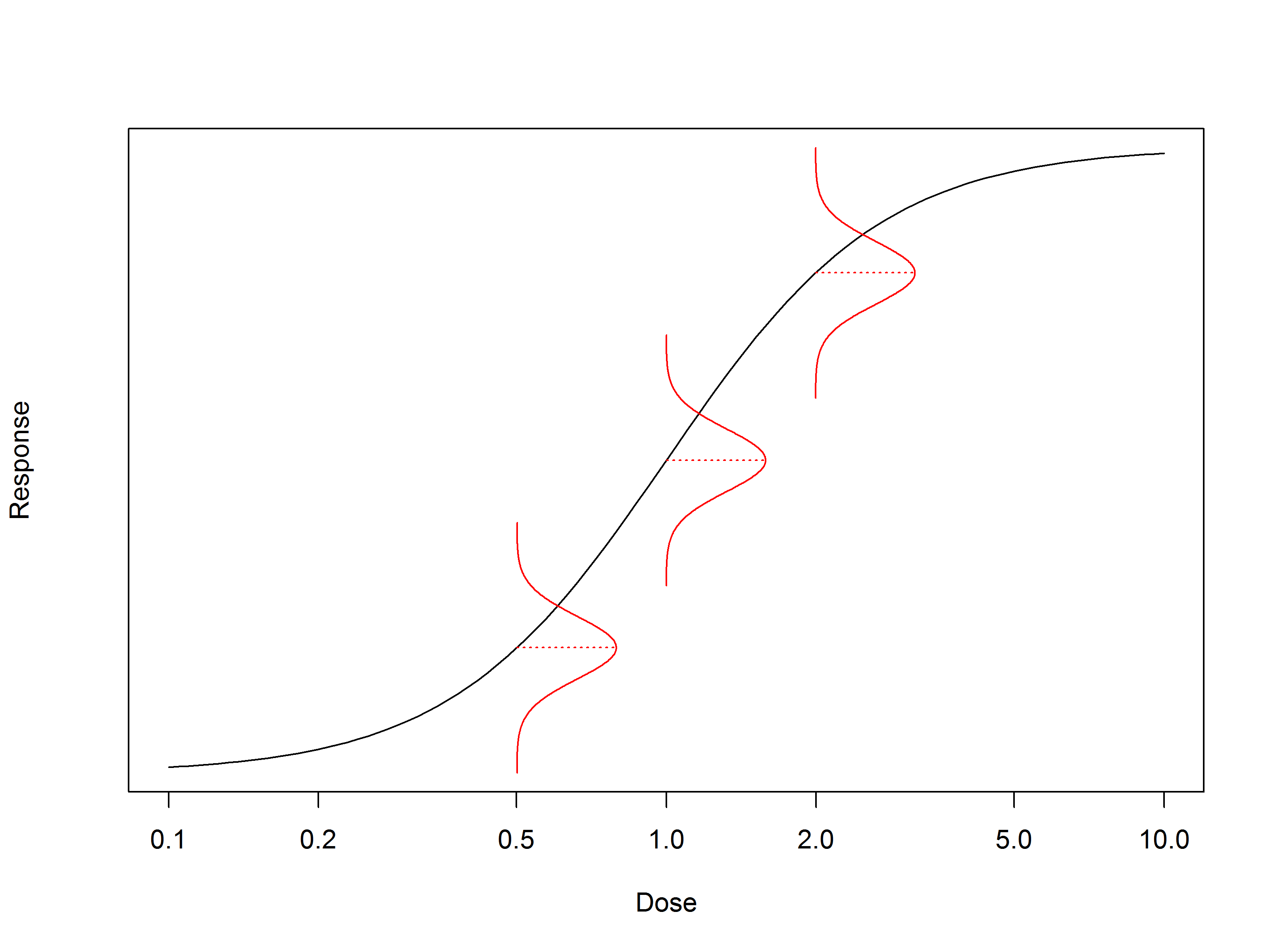
So, let’s start with the simple linear “dose response” model. Actually, this is linear only when we consider response against log (dose).
So if \(y =\) response, \(a =\) intercept and \(x = \log(\text{dose})\): \(y = ax + b\) — a simple linear model.
If we wanted to express this using the raw dose – let’s label this \(z\) – we get \(y = A + B \log(z)\).
Note that the symbols used are changed to reflect the changed definitions.
In this case to bridge between these two we use:
\(A = a\)
\(B = b\)
4 Parameter Logistic
The same goes for the 4PL but it is a bit more complicated! We have a blog dedicated to different parameterisations of the 4PL, so this is only a brief overview here.
The PhEur 5.3 section 3.4 has:
\[u=\delta+\frac{\alpha-\delta}{1+e^{-\beta\left(x-y\right)}}\]
While the USP has:
\[y=D+\frac{A-D}{1+\left(\frac{z}{C}\right)^B}\]
The difference is that the PhEur expresses the formula in terms of \(x = \ln(\text{dose})\), whilst the USP expresses the formula in terms of \(z = \text{dose}\).
So the “bridging” is simple algebra:
\[ \begin{align} x &= \ln(z)\\ \delta &= D\\ \alpha &= A\\ \beta &= -B\\ y &= \ln(C) \end{align} \]
And there you have it – no bridging experiment is required. All you need is a mathematician to do the algebra. Once you have this it is a simple matter to convert all your suitability criteria to the new parameterisation, update the SOP, put the new values in the template, and you are done. Time for tea and brownies…
Whatever the parameterisation, there is zero impact on the model fit, the RP or its confidence interval.
During the presentations, some time was spent trying to relate the sign of the hill slope (B) to the slope as usually described for inhibition and absorption assays. Again this is just maths, there is no need to guess or try data sets to work this out, just differentiate the formulae and you get:
PhEur parameterisation: \(Slope_{\text{PhEur}}=\frac{\left(\alpha-\delta\right)\beta}{4}\)
USP parameterisation: \(Slope_{\text{USP}}=\frac{\left(D-A\right)B}{4}\)
It is now easy to work out whether the \(D\) and \(A\) are left, right, top, or bottom. So, for example, it was pointed out that Gen5 software (which uses the USP parameterisation) fixes \(B\) to be positive. Using the slope equation above for USP, this will mean that for a positive slope assay \((D-A)\) must be positive so \(D\) is the upper asymptote and \(A\) the lower. For a negative slope assay it will be the other way round.
Conclusion
Some of the “different” results presented at BEBPA are entirely predictable and just a matter of maths. Bridging can be done as an exact transfer so there is no impact on validation of the assay.
However, in practise some differences in results are seen despite this direct mathematical conversion. So why is this?
The answer is that it is to do with the algorithms used to calculate the parameters for the curve fit and this is the subject of the next mini blog.